Taxonomic Dereplication vs. Structure-Based Design: Choosing the Right Strategy for Modern Drug Discovery
This article provides a comprehensive comparative analysis of two dominant strategies in natural product and drug discovery: taxonomic-focused dereplication and structure-based approaches.
Taxonomic Dereplication vs. Structure-Based Design: Choosing the Right Strategy for Modern Drug Discovery
Abstract
This article provides a comprehensive comparative analysis of two dominant strategies in natural product and drug discovery: taxonomic-focused dereplication and structure-based approaches. For researchers and drug development professionals, we explore the foundational principles, core methodologies, and practical applications of each paradigm. We detail how taxonomic dereplication, powered by molecular networking and mass spectrometry, enables rapid known-compound filtering to prioritize novelty[citation:2][citation:8]. In contrast, we examine structure-based methods, including virtual screening and molecular docking, which leverage target protein architecture to rationally design or discover bioactive leads[citation:1][citation:6]. The article directly compares their strengths in troubleshooting common pitfalls like rediscovery and off-target effects, and validates their performance through real-world applications in identifying anticancer agents and novel microbial metabolites[citation:1][citation:3]. Finally, we synthesize key decision-making criteria for project-specific strategy selection and outline the future of integrated, AI-enhanced workflows that promise to bridge these complementary philosophies[citation:4][citation:6][citation:7].
Core Philosophies: Defining Taxonomic Prioritization and Structure-Based Rational Design
In the search for new therapeutic agents and the understanding of complex biological systems, researchers are fundamentally guided by a central dichotomy: the identification of the known and the discovery of the unknown. This dichotomy is operationalized through two complementary methodological paradigms: dereplication and de novo discovery. Dereplication is the efficient process of identifying known compounds or taxa within a complex sample to avoid redundant rediscovery, thereby streamlining resource allocation [1] [2]. In contrast, de novo discovery aims to isolate, characterize, and identify entirely novel entities—be they chemical structures, microbial species, or genetic pathways—that are absent from existing databases [3] [4].
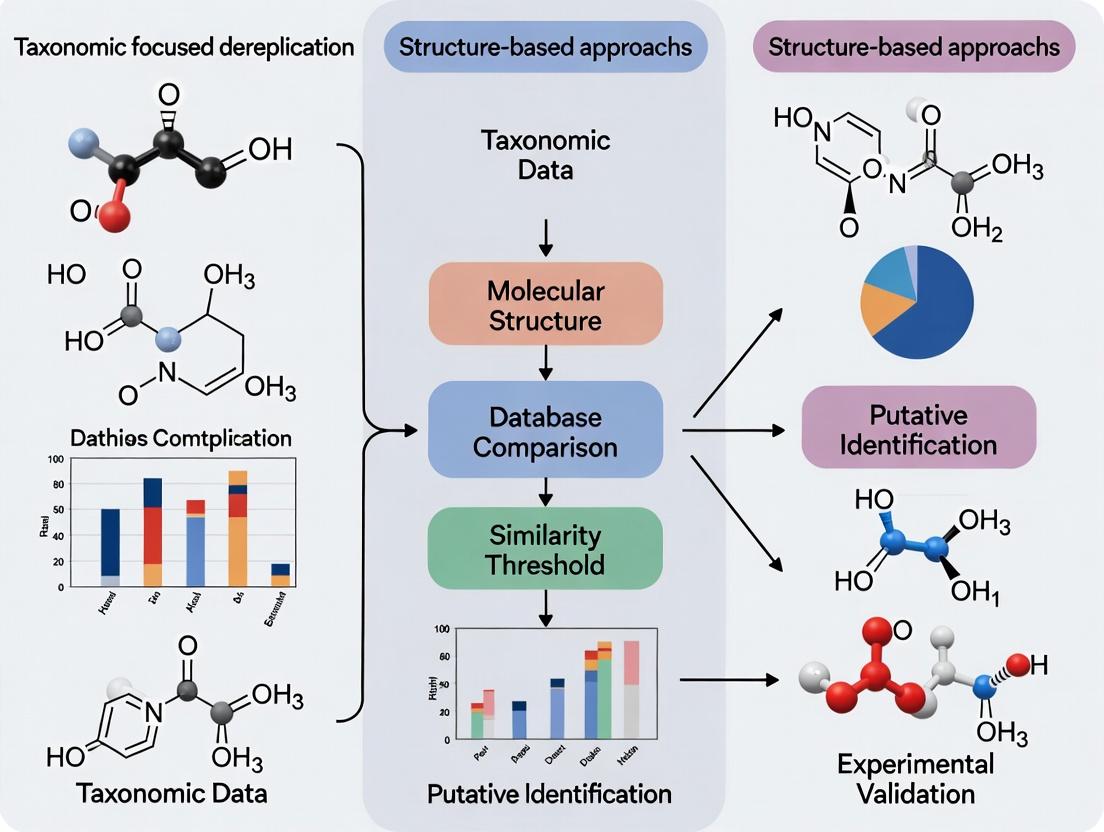
These approaches are framed within two distinct but increasingly convergent research strategies: taxonomic-focused analysis and structure-based approaches. Taxonomic-focused dereplication, prevalent in microbiome research and natural product discovery from biological sources, classifies entities based on evolutionary relationships and marker genes [3] [5]. Structure-based approaches, central to modern drug design, prioritize the three-dimensional architecture and physico-chemical properties of molecular targets and their ligands [6] [7]. This guide provides a comparative analysis of these methodologies, supported by experimental data and protocols, to inform strategic decisions in research and development.
Comparative Analysis: Dereplication vs. De Novo Discovery
In Natural Products & Microbiome Research
This domain leverages analytical chemistry and genomics to navigate the complexity of biological extracts and microbial communities.
Dereplication is a critical first pass to filter out known compounds. As demonstrated in the analysis of a polyherbal liquid formulation, an LC-MS/MS dereplication strategy successfully identified 70 compounds, with 44 uniquely attributed to specific plant species [2]. This process prevents the costly and time-consuming isolation of common metabolites. Similarly, in microbiology, alignment-based (AL) methods map sequencing reads to reference databases (e.g., GTDB, CHOCOPhlAn) to rapidly profile the known taxonomic composition of a sample [3].
De novo discovery targets the uncharted fraction. In soil microbiome research, the use of microbial diffusion chambers enabled the cultivation of previously "uncultivable" bacteria, yielding 1,218 isolates where 16% showed antibiotic activity [4]. This is the experimental counterpart to de novo (DN) bioinformatic approaches, which assemble genomes from sequence data without reference bias, enabling the discovery of novel microbial taxa and gene clusters [3]. An integrated pipeline combining cultivation, bioassay, mass spectrometry (MS) dereplication, and genome mining is optimal for novel natural product discovery [4].
Table 1: Comparison of Approaches in Natural Products & Microbiome Research
| Aspect | Dereplication (Known-First) | De Novo Discovery (Novelty-First) |
|---|---|---|
| Primary Goal | Rapid identification of known entities to avoid rediscovery. | Discovery and characterization of novel entities. |
| Core Methodology | LC-MS/MS spectral matching [1] [2]; Alignment to reference genomic databases [3]. | Bioassay-guided fractionation; Cultivation innovations (e.g., diffusion chambers) [4]; De novo genome assembly & binning [3]. |
| Key Tool/Platform | In-house or public spectral libraries (e.g., GNPS, MassBank) [1]; MetaPhlAn, HUMAnN [3]. | Global Natural Products Social Molecular Networking (GNPS) [8]; Metagenome-Assembled Genome (MAG) reconstruction pipelines. |
| Typical Output | List of annotated compounds or taxa with relative abundances. | Novel chemical structures; Novel microbial genomes & biosynthetic gene clusters (BGCs). |
| Strengths | High speed, efficiency, and reproducibility. Essential for quality control and standardization [2]. | Accesses untapped chemical and biological diversity. Potential for high-impact discovery. |
| Limitations | Limited by scope and quality of reference databases. Blind to novelty. | Resource-intensive, time-consuming, and often low-throughput. |
In Computational Structure-Based Drug Discovery (SBDD)
Here, the dichotomy manifests in the use of known structural information to predict new interactions or to generate novel molecular entities.
Dereplication in SBDD involves screening virtual or chemical libraries against a target to identify known binders or chemotypes. It relies heavily on knowledge-based methods (e.g., machine learning models trained on known protein-ligand complexes) that excel at interpolating within existing chemical space but struggle to generalize to novel scaffolds [6]. The goal is to quickly prioritize compounds with a higher probability of activity based on historical data.
De novo discovery in SBDD refers to the ab initio design or identification of novel molecular scaffolds that optimally fit a target binding site. This is the domain of physics-based methods like molecular docking, free energy perturbation (FEP) calculations, and de novo ligand design algorithms [6] [9]. These methods use principles of molecular mechanics and thermodynamics to evaluate interactions, potentially generating innovative solutions not present in training data.
Table 2: Comparison of Approaches in Computational Structure-Based Drug Discovery
| Aspect | Knowledge-Based (Dereplication-Oriented) | Physics-Based (De Novo-Oriented) |
|---|---|---|
| Primary Goal | Predict activity/affinity by learning from known data. | Predict binding pose and affinity from first physical principles. |
| Core Methodology | Machine Learning (ML) / Deep Learning on structural and bioactivity databases (e.g., PDBbind, ChEMBL) [6]. | Molecular Docking, Molecular Dynamics (MD), Free Energy Perturbation (FEP) [6] [9]. |
| Data Dependency | High; requires large, high-quality training datasets. Performance degrades for novel targets or chemotypes [6]. | Low in principle; but accuracy depends on force-field quality and sampling. Requires a high-resolution target structure. |
| Strength | Extremely fast screening of ultra-large libraries. Excellent for targets rich in data [6]. | Can handle novel scaffolds and make predictions where no ligand data exists. Provides mechanistic insight. |
| Limitation | Risk of overfitting; limited generalizability "outside the box" of training data [6]. | Computationally expensive; can be prone to scoring function inaccuracies; sensitive to input structure quality [6]. |
| Ideal Use Case | Early-stage virtual screening to filter known chemotypes. Lead optimization for data-rich targets. | Hit identification for novel targets. Scaffold hopping and lead optimization for precise affinity prediction. |
Experimental Protocols
This protocol is designed for the rapid identification of known bioactive compounds in complex plant extracts.
- Sample Preparation & Pooling: Prepare standard solutions of reference compounds. Use a log P-based pooling strategy to minimize co-elution and isomer interference. For crude extracts, employ Solid-Phase Extraction (SPE) with C-18 cartridges to remove sugars and interfering matrix components [2].
- LC-MS/MS Analysis: Analyze pools or samples using Reversed-Phase Liquid Chromatography coupled to high-resolution tandem Mass Spectrometry (LC-HRMS/MS). Optimize chromatography for compound separation.
- Data Acquisition: Acquire MS/MS spectra in data-dependent acquisition (DDA) mode. For each compound, collect fragmentation data at multiple collision energies (e.g., 10, 20, 30, 40 eV) and for different adducts ([M+H]⁺, [M+Na]⁺).
- Library Construction & Searching: Construct an in-house spectral library by cataloguing the retention time (RT), precursor m/z, and fragmentation patterns of reference standards. Search experimental MS/MS data against this library and public databases (e.g., GNPS, MassBank). Use a mass error tolerance of <5 ppm for confident annotation [1].
- Validation: Confirm identifications by comparing RT and MS/MS spectra with authentic standards analyzed under identical conditions.
This multi-omic protocol aims to discover novel antibiotics from uncultivated soil bacteria.
- In Situ Cultivation: Construct microbial diffusion chambers with 0.03 µm semi-permeable membranes. Inoculate chambers with a diluted soil slurry in low-nutrient agar and incubate them buried in the source soil for 2-4 weeks to allow growth of uncultivable bacteria.
- Strain Recovery & Screening: Retrieve agar plugs, domesticate isolates on R2A agar, and cryopreserve pure cultures. Screen isolates for bioactivity against target pathogens (e.g., S. aureus, E. coli) using agar overlay assays.
- MS-Based Dereplication: Culture bioactive strains and extract secondary metabolites. Analyze extracts via LC-HRMS/MS. Process data through the GNPS platform for molecular networking and dereplication against spectral libraries to identify known antibiotics.
- Genomic Analysis: Sequence the genome of prioritized bioactive strains. Perform genome mining using tools like antiSMASH to identify Biosynthetic Gene Clusters (BGCs) responsible for secondary metabolite production. This step can reveal the potential for novel compounds even if MS dereplication was unsuccessful.
- Targeted Isolation & Characterization: For strains with novel BGCs or ambiguous MS IDs, scale up fermentation, and use bioassay-guided fractionation to isolate the active compound(s). Elucidate structure using NMR and HRMS.
Visualizing Workflows and Relationships
Integrated De Novo Discovery Workflow
Structure-Based Drug Design Pathways
The Researcher's Toolkit
Table 3: Essential Reagents and Materials for Featured Experiments
| Item | Function / Application | Relevant Protocol |
|---|---|---|
| 0.03 µm Polycarbonate Membrane | Forms the semi-permeable barrier of diffusion chambers, allowing nutrient exchange while containing microorganisms [4]. | De Novo Antibiotic Discovery |
| R2A Agar / SMS Agar | Low-nutrient cultivation media used to recover and grow oligotrophic soil bacteria that fail to grow on rich media [4]. | De Novo Antibiotic Discovery |
| C-18 Solid Phase Extraction (SPE) Cartridge | Removes polar interfering substances (e.g., sugars, salts) from complex herbal extracts, reducing matrix effects and improving LC-MS signal clarity [2]. | LC-MS/MS Dereplication |
| LC-MS Grade Solvents (MeOH, H₂O with Formic Acid) | High-purity mobile phase for liquid chromatography to ensure reproducible retention times and prevent ion source contamination in MS [1] [2]. | Both Protocols |
| Authentic Chemical Standards | Reference compounds used to build in-house spectral libraries for definitive identification by matching retention time and MS/MS spectrum [1]. | LC-MS/MS Dereplication |
| Syto9 / DAPI Nucleic Acid Stains | Fluorescent dyes used to count microbial cells in soil slurries for standardized inoculation of diffusion chambers [4]. | De Novo Antibiotic Discovery |
| Global Natural Products Social (GNPS) Platform | A public online platform for sharing and analyzing mass spectrometry data, enabling spectral library matching and molecular networking [4] [8]. | Both Protocols |
The dichotomy between dereplication and de novo discovery is not a barrier but a strategic framework. The most effective research pipelines in both natural products and computational drug discovery are those that sequentially integrate both paradigms. The future lies in hybrid approaches: using dereplication to efficiently clear the known landscape, thereby focusing costly de novo efforts on the most promising unexplored territories [3] [4]. Similarly, in SBDD, combining the speed of knowledge-based methods with the rigorous, generative potential of physics-based simulations represents the state of the art [6] [7]. Whether focused on taxonomy or molecular structure, the ultimate goal remains the same: to navigate the vast universe of the unknown by first intelligently managing the known.
The discovery of new therapeutic agents has undergone a fundamental paradigm shift, evolving from observation-driven natural product isolation to prediction-enabled molecular design. This transition represents more than a mere technological upgrade; it signifies a profound change in the philosophical approach to interrogating biological systems and chemical space. Historically, bioactivity-guided fractionation served as the cornerstone of drug discovery, relying on systematic biological screening of complex natural extracts to isolate active compounds, often informed by traditional medicinal knowledge [10]. In parallel, the natural products field developed taxonomy-focused dereplication—a strategy to efficiently identify known compounds from biological sources based on taxonomic relationships and spectroscopic data, thereby avoiding redundant rediscovery [11].
Conversely, the rise of computational first principles and structure-based approaches has introduced a target-centric, rational framework. Enabled by advances in structural biology, high-performance computing, and machine learning, this paradigm uses the three-dimensional structure of therapeutic targets to design or discover ligands with precision [12] [13]. This guide provides an objective comparison of these foundational methodologies, examining their performance, experimental requirements, and ideal applications within modern drug development. The analysis is framed by the broader research thesis that contrasts the organism- and chemistry-centric viewpoint of taxonomy-focused dereplication with the target- and structure-centric viewpoint of computational design, highlighting how their integration is shaping the future of the field.
Performance Comparison: Key Metrics and Outcomes
The following tables quantitatively compare the core characteristics, outputs, and practical performance of taxonomy-focused dereplication and structure-based computational approaches.
Table 1: Foundational Characteristics and Strategic Focus
| Comparison Aspect | Taxonomy-Focused Dereplication & Bioactivity-Guided Fractionation | Computational First Principles & Structure-Based Design |
|---|---|---|
| Primary Objective | Identify novel bioactive compounds from nature; avoid re-isolation of knowns [11] [10]. | Design or discover novel ligands for a defined macromolecular target [12] [13]. |
| Starting Point | Biological material (plant, microbial extract) with observed bioactivity or taxonomic lineage [10]. | 3D structure of a target protein (experimental or predicted) [12]. |
| Core Principle | Leverage evolutionary conservation of biosynthetic pathways within taxa for targeted discovery [11]. | Apply principles of molecular recognition, thermodynamics, and docking physics [13] [6]. |
| Key Data Inputs | Taxonomic classification; LC-MS/MS, NMR spectroscopic data [11] [14]. | Protein atomic coordinates; chemical libraries; force field parameters [12] [6]. |
| Typical Output | Isolated and characterized novel natural product(s) with confirmed biological activity [10]. | Predicted high-affinity small molecule binders with a proposed binding pose [13]. |
Table 2: Quantitative Performance and Practical Metrics
| Performance Metric | Taxonomy-Focused/Bioactivity-Guided Approach | Computational/Structure-Based Approach | Supporting Data & Context |
|---|---|---|---|
| Historical Success (FDA-Approved Drugs) | A major source: ~35% of modern medicines are natural products or direct derivatives [10]. | Significant contributor: SBDD estimated to have contributed to >200 approved drugs; FBDD directly led to 4 [15]. | Natural products dominate in anti-infectives and oncology [10]. SBDD is versatile across target classes [12]. |
| Development Timeline (Early Stage) | Can be lengthy due to slow extraction, fractionation, and structure elucidation steps [10]. | Rapid virtual screening of ultra-large libraries (billions of compounds) is possible in weeks [13]. | Computational speed is offset by later synthetic and experimental validation requirements. |
| Cost Implications | High costs associated with large-scale biomass collection, purification, and wet-lab screening [10]. | Can reduce early discovery costs significantly; CADD estimated to cut discovery costs by up to 50% [13]. | Major costs in the natural product pipeline are front-loaded; computational costs are largely in infrastructure/software. |
| Hit Rate Efficiency | Low hit rate in random screening; greatly enhanced by ethnopharmacological or taxonomic focus [10]. | Typical experimental hit rates from structure-based virtual screening range from 10% to 40% [13]. | Hit rate for computational methods depends heavily on target "druggability" and library quality [6]. |
| Chemical Space Coverage | Explores biologically pre-validated, structurally complex, and often "drug-like" chemical space [14]. | Can theoretically access vast synthetic chemical space (e.g., >6.7 billion in REAL database) [13]. | Natural products cover regions of chemical space often inaccessible to synthetic libraries [14]. |
| Major Challenge | Supply, re-isolation of knowns, slow dereplication, complex structure determination [11] [10]. | Accurate scoring and affinity prediction, handling protein flexibility, synthetic accessibility of hits [13] [6]. | Both fields are developing solutions: genomic mining for NPs; better force fields & AI for SBDD [6] [14]. |
Experimental Protocols: Representative Methodologies
Protocol for Taxonomy-Focused Dereplication Using 13C NMR
This protocol, based on the CNMR_Predict pipeline, creates a targeted database for efficient dereplication of natural products from a specific organism [11].
1. Taxon Definition and Raw Data Acquisition:
- Select the organism of interest (e.g., Brassica rapa subsp. rapa L.).
- Query the LOTUS database (or similar taxonomy-aware NP database) using the species name as a keyword.
- Download all associated chemical structures in a standard file format (e.g., SDF V3000).
2. Data Curation and Standardization:
- Remove duplicate structures based on canonical identifiers (e.g., InChI keys).
- Apply tautomer normalization to ensure all structures are in a consistent, predictable form (e.g., converting iminol forms to amides).
- Adjust atomic valence representations to ensure compatibility with downstream prediction software.
3. Spectral Data Prediction and Database Creation:
- Import the curated structure list into specialized 13C NMR prediction software (e.g., ACD/Labs CNMR Predictor).
- Execute batch prediction of chemical shifts for all carbon atoms in all compounds.
- Export the combined dataset (structures + predicted spectra) into a searchable, taxon-specific database.
4. Experimental Dereplication:
- Obtain a 13C NMR spectrum (or a 1D 1H spectrum with 13C projections) of the purified unknown compound or complex mixture.
- Search the experimental chemical shifts against the created taxon-specific database.
- Evaluate matches based on chemical shift tolerance (e.g., ± 0.5-1.0 ppm) and the number of matching signals. A high-confidence match indicates a known compound, halting further costly isolation efforts.
Protocol for High-Throughput X-ray Crystallography Fragment Screening (FBDD)
This protocol outlines a primary fragment screening approach using high-throughput X-ray crystallography, as implemented at facilities like XChem [15].
1. Target and Library Preparation:
- Protein Target: Produce and purify a stable, crystallizable target protein (>10 mg, >95% pure). Engineer constructs if necessary for crystallization.
- Fragment Library: Obtain a curated library of 500-1500 fragments compliant with the "Rule of Three" (MW <300, cLogP ≤3, ≤3 H-bond donors/acceptors, etc.) [15].
- Co-crystallization or Soaking: Prepare high-quality, reproducible protein crystals. For screening, use a fragment soaking method:
- Group fragments into cocktails of 4-8 compounds, each at high concentration (e.g., 200 mM in DMSO).
- Briefly soak crystals in mother liquor containing the fragment cocktail.
- Alternatively, perform co-crystallization with individual fragments or cocktails.
2. High-Throughput Data Collection and Processing:
- Harvest and cryo-cool soaked crystals.
- Collect X-ray diffraction data for hundreds to thousands of crystals in an automated, high-throughput beamline setup.
- Process data automatically: integrate diffraction images, scale intensities, and solve structures by molecular replacement using the native protein model.
3. Hit Identification and Analysis:
- Use automated electron density analysis software (e.g., PanDDA) to detect bound fragments in the difference electron density maps, even for weak binders [15].
- Manually validate hits: inspect electron density for clear fragment shape, assess fit, and model the fragment into the density.
- Output: A list of confirmed fragment hits with precise, experimentally determined binding modes, locations (orthosteric/allosteric), and protein-ligand interaction maps.
4. Hit-to-Lead Progression:
- Prioritize hits based on binding pose, ligand efficiency, and chemical tractability.
- Initiate fragment growing, linking, or optimization using the detailed structural information to design more potent lead compounds [15].
Workflow and Relationship Diagrams
Diagram 1: Bioactivity-Guided Fractionation with Dereplication Workflow (Max Width: 760px).
Diagram 2: Computational First Principles Drug Discovery Workflow (Max Width: 760px).
Table 3: Key Reagents and Resources for Dereplication and Structure-Based Research
| Tool/Resource Name | Category | Primary Function | Relevant Paradigm |
|---|---|---|---|
| LOTUS Database (lotus.naturalproducts.net) | Database | Provides rigorously curated links between natural product structures and their taxonomic sources for targeted queries [11]. | Taxonomy-Focused Dereplication |
| ACD/Labs CNMR Predictor and DB | Software | Predicts 13C NMR chemical shifts from molecular structure to create searchable spectral databases for dereplication [11]. | Taxonomy-Focused Dereplication |
| GNPS (Global Natural Products Social Molecular Networking) | Data Platform | Enables community-wide sharing and curation of MS/MS spectral data for annotation and dereplication of natural products [14]. | Bioactivity-Guided Fractionation |
| Rule of Three (Ro3) Fragment Library | Chemical Library | A curated collection of small, simple molecules (MW <300) used to probe protein binding sites and identify weak but efficient starting points for drug design [15]. | Fragment-Based Drug Discovery (FBDD) |
| XChem/High-Throughput X-ray Crystallography Platform | Experimental Platform | Enables primary screening of fragment libraries by obtaining protein-ligand co-crystal structures at scale, providing direct structural data on binding [15]. | FBDD / Structure-Based Design |
| AlphaFold Protein Structure Database | Database/Algorithm | Provides highly accurate predicted protein 3D models for targets with no experimental structure, massively expanding the scope of SBDD [13]. | Computational First Principles |
| Enamine REAL (REadily AccessibLe) Database | Chemical Library | An ultra-large, commercially available virtual library of synthesizable compounds (>6.7 billion) for virtual screening [13]. | Structure-Based Virtual Screening |
| Relaxed Complex Method | Computational Method | Uses conformational ensembles from Molecular Dynamics simulations for docking, accounting for protein flexibility and cryptic pockets [13]. | Dynamics-Based Drug Discovery |
Discussion and Concluding Comparison
The contrast between taxonomy-focused dereplication and computational first-principles design encapsulates a broader evolution in drug discovery: from observation and exploitation of nature's chemical bounty to prediction and engineering of molecular interactions. The former approach, rooted in biology and chemistry, excels at delivering structurally novel, biologically pre-validated scaffolds that have historically been a major source of drugs, especially in challenging areas like oncology and infection [10] [14]. Its strengths lie in its connection to biologically relevant chemical space and its clear path to a bioactive compound. Its primary weaknesses are throughput, scalability, and the inherent uncertainty of the discovery process.
The latter approach, rooted in physics and computer science, offers speed, scalability, and rational design. It can rapidly explore vast synthetic chemical spaces, provide atomic-level insight into mechanism, and systematically optimize compounds. Its success is evident in the hundreds of approved drugs it has contributed to [15] [12]. Its critical limitations revolve around the accuracy of scoring functions, the challenges of modeling flexible biological systems, and the ultimate need to synthesize and test predicted compounds in the lab [13] [6].
The forward-looking thesis of modern drug discovery is not the supremacy of one paradigm over the other, but their strategic integration. Computational methods are revolutionizing natural product research through genome mining, spectral prediction, and database dereplication [11] [14]. Conversely, natural product-derived scaffolds are inspiring the design of focused libraries for virtual screening [10]. The most powerful future workflows will likely be hybrid: using taxonomic and genomic intelligence to guide the selection of natural sources, applying advanced analytics and dereplication to quickly identify novel chemotypes, and then using structure-based design to optimize these natural scaffolds into potent, drug-like candidates. This synthesis of biological wisdom and computational power represents the next chapter in the historical context of therapeutic discovery.
The process of dereplication—the rapid identification of known compounds in complex mixtures to prioritize novel entities—is a critical bottleneck in natural product discovery and microbiome analysis. Traditionally, approaches have been bifurcated into structure-based methods, which prioritize chemical features, and taxonomic-focused methods, which leverage the evolutionary relationships of the source organism [16] [11]. This guide objectively compares the performance of taxonomic-focused dereplication, which integrates phylogeny and spectral libraries, against alternative structure-based and similarity-based approaches.
The core thesis posits that taxonomic-focused dereplication provides a more efficient framework for annotating known compounds and predicting novel chemical space by constraining identification within evolutionary boundaries. This approach is particularly powerful when coupled with public spectral libraries and phylogenetic placement algorithms, enabling researchers to bypass the re-isolation of known compounds and accelerate the discovery pipeline [17] [18].
Core Concepts and Comparative Frameworks
Taxonomic-focused dereplication operates on the principle that biosynthetic pathways are often conserved within taxonomic groups (e.g., genera, families). By knowing the source organism's phylogeny, the search space for compound identification is significantly reduced. This method commonly utilizes phylogenetic placement of marker genes or whole genomes, alongside targeted or non-targeted mass spectrometry, to identify compounds [19] [18].
In contrast, structure-based dereplication is agnostic to taxonomy. It relies on direct comparison of analytical data—such as mass spectra, NMR shifts, or fragmentation patterns—against comprehensive libraries of pure compound data. The identification is based purely on spectral similarity, often using computational tools like molecular networking [17] [16].
A third paradigm, prominent in metagenomics, is the alignment-based (AL) vs. de novo (DN) approach. AL methods map sequencing reads to reference databases for rapid profiling of known taxa, while DN methods assemble reads without references to discover novel genomic elements [20] [21]. The choice between these strategies presents a trade-off between speed, reliance on existing knowledge, and the ability to discover novelty.
Table 1: Core Conceptual Comparison of Dereplication Strategies
| Strategy | Primary Data Input | Key Mechanism | Main Advantage | Primary Limitation |
|---|---|---|---|---|
| Taxonomic-Focused | Genetic material (DNA/RNA) & Spectral Data | Phylogenetic placement & taxonomic spectral filtering | Constrains search space, predicts novel related compounds | Requires phylogenetic knowledge; limited by reference databases |
| Structure-Based | Spectroscopic data (MS, NMR) | Direct spectral matching & molecular networking | Taxonomy-agnostic; direct compound-level identification | Prone to ambiguous matches for isomers; overlooks taxon-specific novelty |
| Alignment-Based (AL) | Sequencing reads (e.g., shotgun) | Mapping to reference genomes/marker genes | Fast, efficient for profiling known communities | Biased against novel taxa not in reference databases [20] |
| De Novo (DN) | Sequencing reads (e.g., shotgun) | De novo assembly and binning | Discovers novel taxa and genes; reference-independent | Computationally intensive; higher data sparsity [20] [21] |
Performance Comparison: Experimental Data and Outcomes
Spectral Library Development and Application
The development of specialized, public spectral libraries exemplifies the power of focused resources. The Pyrrolizidine Alkaloid Spectral Library (PASL) contains 165 MS/MS spectra from 102 compounds (84 standards, 18 from crude extracts). When applied to dereplicate compounds in plant extracts, this taxonomy-focused library (targeting Asteraceae, Boraginaceae, Fabaceae) enabled rapid annotation without pure standards. In a comparative sense, a broader, untargeted search of generic MS/MS libraries would yield higher rates of false annotations due to the structural diversity across all plant taxa [17].
Metagenomic Profiling: Alignment vs. De Novo
A direct comparison of AL and DN methods on the same gut microbiome dataset (346 samples) reveals a clear trade-off [20]:
- AL Methods (e.g., MetaPhlAn) produced taxonomic profiles with lower sparsity and identified a greater number of differentially abundant taxa associated with host BMI. They explained a greater proportion of variance (~8.7%).
- DN Methods (MAG reconstruction) captured more novel microbial diversity (e.g., higher relative abundance of Archaea) but produced sparser matrices, identifying only a subset of the significant taxa found by AL. However, DN enabled functional insights, such as identifying a novel 2,5-diketo-D-gluconate reductase A enzyme in Alistipes onderdonkii MAGs [20].
An integrative tool like the Read Annotation Tool (RAT), which combines signals from MAGs, contigs, and reads, demonstrates the hybrid performance gain. In benchmark tests on CAMI2 data, RAT achieved superior precision and sensitivity by first inheriting reliable taxonomy from assembled sequences before annotating remaining reads [21].
Table 2: Quantitative Performance Comparison from Key Studies
| Study / Tool | Approach | Key Performance Metric | Result | Implication for Dereplication |
|---|---|---|---|---|
| Pyrrolizidine Alkaloid Library (PASL) [17] | Taxonomy-focused spectral matching | Library scope & application | 102 PAs, 165 MS/MS spectra; successful dereplication in plant extracts | Focused libraries reduce false positives in targeted taxon groups. |
| AL vs. DN Microbiome Analysis [20] | Alignment-based vs. De novo assembly | Number of significant taxa & novelty discovery | AL identified more diff. abundant taxa; DN found novel enzyme genes. | AL is efficient for known communities; DN is essential for functional novelty. |
| RAT (Read Annotation Tool) [21] | Integrative (MAGs + contigs + reads) | Precision & sensitivity on CAMI2 data | Outperformed state-of-the-art profilers by integrating multi-level signals. | Leveraging assembled data significantly improves read annotation accuracy. |
| Phylogenetic Placement [18] | Phylogeny-based sequence placement | Taxonomic assignment accuracy | Provides evolutionary context, increasing accuracy over simple similarity. | Essential for placing novel sequences from poorly characterized taxa. |
Phylogenetic Placement for Taxonomic Assignment
Phylogenetic placement, a cornerstone of taxonomic-focused analysis, directly addresses the limitations of similarity-based (BLAST) searches. By placing query sequences within a fixed reference tree, it considers evolutionary history and branch lengths, leading to more accurate taxonomic assignment, especially for novel or divergent sequences [18]. A review of the first decade of these methods confirms they eliminate the requirement for exact database matches and reduce misidentification, providing a robust framework for analyzing metabarcoding data from diverse environments [18].
Detailed Experimental Protocols
This protocol details the creation of the Pyrrolizidine Alkaloid Spectral Library (PASL).
- Sample Preparation: Dilute pure analytical standards to ~500 μg/L in 10% methanol. For crude extracts, homogenize 10 mg of freeze-dried plant material in 1 mL of 0.2% formic acid, filter (0.45 µm), and analyze directly.
- LC-HRMS/MS Analysis:
- System: UHPLC (e.g., Thermo Vanquish) coupled to an Orbitrap mass spectrometer (e.g., Q Exactive).
- Chromatography: C18 column (2.1 x 150 mm, 1.7 µm). Gradient: 10 mM ammonium carbonate buffer (pH 9) and acetonitrile over 18.5 minutes.
- MS Acquisition: Full-scan MS1 (resolution: 120,000 FWHM) followed by data-dependent MS/MS (resolution: 15,000 FWHM) using stepped collision energies (e.g., 15, 30, 45 eV).
- Data Processing & Library Curation:
- Convert raw files to open
.mzMLformat. - Use workflows (e.g., MSMSChooser on GNPS) to extract clean, representative MS/MS spectra for each compound.
- Annotate compounds with validated structures and isomeric SMILES, correcting for database errors in stereochemistry.
- Validate the library by performing molecular networking against public GNPS libraries and applying it to dereplicate known PAs in test plant extracts.
- Convert raw files to open
This protocol uses assembly-derived signals to improve read annotation.
- Assembly and Binning: Perform de novo assembly of quality-filtered metagenomic reads using a tool like MEGAHIT or metaSPAdes. Bin contigs into Metagenome-Assembled Genomes (MAGs) using tools like MetaBAT2.
- Annotation of Long Sequences:
- Annotate contigs using the Contig Annotation Tool (CAT).
- Annotate MAGs using the Bin Annotation Tool (BAT). Both CAT and BAT predict Open Reading Frames (ORFs), query them against a protein database (e.g., NCBI nr or GTDB) via DIAMOND, and assign taxonomy based on consensus.
- Read Mapping and Inheritance: Map all quality-filtered reads back to the contigs using BWA-MEM. A read inherits the taxonomy of the contig (or MAG) it maps to.
- Direct Annotation of Unmapped Reads: Annotate reads that do not map to contigs, and contigs not annotated by CAT, by directly querying them against the protein database using DIAMOND blastx.
- Profile Integration: Combine the inherited annotations (high reliability) with the direct read annotations (higher coverage) to produce a comprehensive, high-fidelity taxonomic profile.
This protocol outlines creating a custom 13C NMR database for a specific taxon.
- Compound Retrieval: Query the LOTUS database (or other NP database) using the target taxon name (e.g., Brassica rapa) to download associated chemical structures in SDF format.
- Structure Curation: Process the SDF file with scripts to remove duplicates, correct tautomeric forms (e.g., iminol to amide), and standardize valence descriptions for compatibility with prediction software.
- Spectral Prediction: Import the curated SDF file into spectral prediction software (e.g., ACD/Labs CNMR Predictor). Calculate and predict 13C NMR chemical shifts for all structures.
- Database Deployment: Export the combined structural and predicted NMR data into a searchable format. This custom database can then be used to dereplicate mixtures from the target taxon by matching observed 13C NMR chemical shifts.
Visualization of Workflows and Logical Frameworks
Integrated Dereplication Workflow for Natural Products and Microbiomes
Taxonomic vs. Structure-Based Dereplication Strategy Comparison
Table 3: Key Reagents, Tools, and Databases for Taxonomic-Focused Dereplication
| Category | Item / Resource | Specific Example / Vendor | Primary Function in Dereplication |
|---|---|---|---|
| Spectral Libraries | Public MS/MS Libraries | GNPS Mass Spectrometry Libraries [17] | Repository for experimental spectra for spectral matching and networking. |
| Taxon-Focused MS Library | Pyrrolizidine Alkaloid Spectral Library (PASL) [17] | Targeted library for rapid, accurate dereplication within a toxin class. | |
| NMR Prediction Software | ACD/Labs CNMR Predictor [11] | Predicts 13C NMR shifts to build custom taxon-focused databases. | |
| Phylogenetic & Genomic Tools | Phylogenetic Placement Engine | EPA-ng, pplacer [18] | Places query sequences on a reference tree for taxonomy and evolution insight. |
| Profiling & Assembly Tools | MetaPhlAn4 (AL), MEGAHIT (DN) [20] | AL: Rapid taxonomic profiling. DN: De novo metagenomic assembly. | |
| Integrative Annotation Pipeline | CAT/BAT/RAT Pack [21] | Annotates contigs, bins, and reads for comprehensive, accurate profiles. | |
| Databases | Natural Product Database | LOTUS [11] | Links NP structures to taxonomic origin for taxon-focused queries. |
| Taxonomic Reference Database | Genome Taxonomy Database (GTDB) [21] | Standardized microbial taxonomy for robust classification. | |
| General Protein Database | NCBI non-redundant (nr) database [21] | Reference for homology searches in annotation pipelines. | |
| Experimental Materials | Chromatography Column | Waters Acquity UPLC BEH C18 (1.7µm) [17] | High-resolution separation of complex extracts prior to MS analysis. |
| Internal Standard (for quant.) | Deuterated or analog compounds (e.g., Heliotrine) [17] | Ensures quantitative reliability and reproducibility in LC-MS. |
The pursuit of new therapeutic agents stands at a crossroads between two fundamental philosophies: the taxonomy-focused dereplication of known chemical entities and the de novo structure-based design of novel compounds [11] [22]. Dereplication efficiently identifies known compounds within complex natural extracts, preventing redundant research by leveraging databases linked to biological taxonomy and spectroscopic data [11]. In contrast, structure-based drug discovery (SBDD) utilizes the three-dimensional architecture of a biological target to rationally design or discover novel ligands that modulate its function [6] [23].
This guide focuses on the computational core of SBDD, charting the evolution from rapid, static docking methods to sophisticated, dynamic simulations. Molecular docking provides a crucial first pass, predicting how a small molecule might fit into a protein's binding site [24] [25]. However, this static snapshot often fails to capture the dynamic reality of biomolecular recognition. Molecular dynamics (MD) simulations address this by modeling the physical movements of atoms over time, offering insights into conformational changes, binding pathways, and binding stability [24] [25]. The field is now being revolutionized by artificial intelligence and deep learning models that predict ligand-specific protein conformational changes, heralding a new era of "dynamic docking" [26] [27].
The integration of these computational tiers—from fast screening to high-fidelity simulation—creates a powerful pipeline. This pipeline is increasingly augmented by experimental structural biology techniques like cryo-EM and solution-state NMR, which provide critical high-resolution data and validate computational predictions [6] [28]. This guide will objectively compare the performance, data requirements, and optimal applications of these approaches, providing researchers with a framework for method selection within the broader drug discovery landscape.
Comparative Performance Analysis: Static Docking vs. Dynamic Simulations
The choice between static docking and dynamic simulation is governed by a trade-off between computational speed and biological fidelity. The table below summarizes their core performance characteristics, supported by benchmark data and practical applications.
Table 1: Performance Comparison of Static Docking and Dynamic Simulation Approaches
| Aspect | Static Molecular Docking | Molecular Dynamics (MD) Simulation | AI-Driven Dynamic Docking (e.g., DynamicBind) [27] |
|---|---|---|---|
| Primary Objective | Predict optimal binding pose and rank ligands by affinity [24] [25]. | Model time-dependent behavior, stability, and conformational changes of the complex [24] [25]. | Predict ligand-specific protein conformations and poses from apo structures [27]. |
| Timescale | Seconds to minutes per ligand [24]. | Nanoseconds to microseconds, requiring days to weeks of compute time [24]. | Minutes to hours per ligand on GPU hardware [27]. |
| Treatment of Flexibility | Limited; typically fully flexible ligand with a rigid or semi-flexible (side-chains only) receptor [25]. | Full atomic flexibility for both ligand and receptor, including solvent [24] [25]. | Models large-scale backbone and side-chain conformational changes driven by the ligand [27]. |
| Key Output Metrics | Docking score (kcal/mol), predicted binding pose, interaction maps [24]. | Trajectory files, RMSD/RMSF, hydrogen bond occupancy, free energy of binding (ΔG) [24]. | Predicted ligand pose RMSD, protein pocket RMSD (vs. holo structure), clash scores [27]. |
| Typical Application | High-throughput virtual screening of 1,000 - 10^6 compounds [24] [25]. | Detailed mechanistic study, binding stability validation, and lead optimization for a few candidates [24] [29]. | Pose prediction and virtual screening where large receptor flexibility or cryptic pockets are involved [27]. |
| Pose Prediction Accuracy (RMSD < 2Å) | Varies (20-70%) highly dependent on target and software; degrades with receptor flexibility [25]. | High for stable binding modes sampled from a correct starting pose; not used for primary screening. | Reported 33-39% success on challenging benchmarks using only AlphaFold-predicted apo structures [27]. |
| Success in Virtual Screening (Enrichment) | Moderate; limited by scoring function accuracy and rigid receptor approximation [6] [25]. | Not directly applicable due to prohibitive cost. | Demonstrates state-of-the-art performance in virtual screening benchmarks [27]. |
| Computational Cost | Very Low | Very High | Moderate |
Supporting Experimental Data: A 2024 study on antiviral discovery provides a direct comparison. Molecular docking screened 200 natural metabolites against viral RNA polymerase, identifying leads like cytochalasin Z8 (docking score: -8.9 kcal/mol). Subsequent 200-ns MD simulations on the top candidates confirmed complex stability, with root-mean-square deviation (RMSD) profiles plateauing, validating the docking-predicted poses [29]. This two-tiered approach is a standard validation protocol.
AI-driven dynamic docking, as exemplified by DynamicBind, addresses a key weakness of static docking. Benchmarking on the PDBbind and Major Drug Target sets showed DynamicBind successfully predicted ligand poses within 2Å RMSD in 33-39% of cases using only AlphaFold-predicted apo structures, outperforming traditional docking tools like GNINA and GLIDE [27]. Its ability to sample large conformational changes (e.g., DFG-in/out transitions in kinases) is particularly notable where static methods fail [27].
Experimental Protocols: From Virtual Screening to Simulation
A robust structure-based workflow typically progresses from broad screening to focused, high-fidelity analysis. Below are detailed protocols for a standard docking-MD validation pipeline and an emerging AI-based dynamic docking approach.
Protocol 1: Integrated Docking and MD Simulation for Lead Validation [29]
Target Preparation:
- Obtain the 3D structure of the target protein from the Protein Data Bank (PDB) or via homology modeling.
- Process the structure: remove water molecules and heteroatoms (except crucial cofactors), add hydrogen atoms, assign protonation states, and repair missing residues/loops.
- For docking, define the binding site using a grid box centered on known catalytic residues or a reference ligand.
Ligand Library Preparation:
- Compile 2D structures (e.g., SMILEs) of compounds from chemical databases (e.g., PubChem, ZINC).
- Generate 3D conformations, minimize energy, and assign appropriate charges and rotatable bonds.
- Convert both protein and ligand files to the required format (e.g., PDBQT for AutoDock).
Molecular Docking Execution:
- Perform docking using software like AutoDock Vina or Glide.
- Set exhaustiveness/search parameters appropriately. Run the docking simulation to generate multiple poses per ligand.
- Rank compounds based on docking score (estimated binding affinity in kcal/mol).
Post-Docking Analysis & Selection:
- Visually inspect top-ranked poses for sensible interaction patterns (hydrogen bonds, hydrophobic contacts).
- Apply drug-likeness filters (e.g., Lipinski's Rule of Five).
- Select the top 1-5 compounds for further dynamic analysis.
Molecular Dynamics Simulation:
- Solvate the top docked complex in an explicit water box (e.g., TIP3P model). Add ions to neutralize the system.
- Energy minimize the system to remove steric clashes.
- Gradually heat the system to 310 K under constant volume (NVT ensemble), then equilibrate at constant pressure (NPT ensemble, 1 atm).
- Run the production MD simulation for a defined time (e.g., 50-200 ns). Use a 2-fs integration time step.
- Analyze trajectories: calculate RMSD (complex stability), RMSF (residue flexibility), hydrogen bond occupancy, and MM/PBSA-based binding free energy.
Protocol 2: AI-Based Dynamic Docking with DynamicBind [27]
Input Preparation:
- Protein Input: Use an apo (unbound) protein structure, preferably an AlphaFold2-predicted model in PDB format.
- Ligand Input: Provide the small molecule in a standard format (SMILES or SDF). The model uses RDKit to generate an initial 3D conformation.
Model Inference:
- The ligand is initially placed randomly around the protein.
- The DynamicBind model, an SE(3)-equivariant geometric diffusion network, executes a series of iterative updates (e.g., 20 steps).
- Initially, it adjusts only the ligand's conformation and placement. Subsequently, it jointly optimizes both ligand pose and protein side-chain (and in part backbone) conformations to reach a complementary, low-energy complex.
Output and Selection:
- The model generates an ensemble of predicted complex structures.
- A built-in scoring module (contact-LDDT or cLDDT) ranks the predictions based on estimated accuracy.
- The top-ranked structure is selected as the final predicted holo-complex, which can be used for virtual screening prioritization or detailed interaction analysis.
Visualization of Workflows and Relationships
Diagram: Parallel Workflows of Dereplication and Structure-Based Design
Diagram: Method Spectrum in Structure-Based Approaches
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Software and Data Resources for Structure-Based Research
| Category | Item Name | Function & Application | Key Characteristics |
|---|---|---|---|
| Docking & Screening | AutoDock Vina [24] [25] | Open-source software for molecular docking and virtual screening. | Uses a gradient-optimized scoring function; fast and widely used in academia. |
| Glide (Schrödinger) [25] [26] | High-accuracy docking software for pose prediction and virtual screening. | Employs systematic search and empirical scoring; a commercial industry standard. | |
| Molecular Dynamics | GROMACS [24] | Open-source, high-performance MD simulation package. | Extremely fast for biomolecular systems; runs on CPUs and GPUs. |
| AMBER [24] | Suite of MD simulation programs with specialized force fields. | Includes sophisticated tools for free energy calculation (MM/PBSA/GBSA). | |
| AI & Advanced Modeling | DynamicBind [27] | Deep learning model for dynamic docking and ligand-induced conformational prediction. | Predicts holo-like complexes from apo structures; handles large conformational changes. |
| AlphaFold2 [6] [27] | Deep learning system for highly accurate protein structure prediction. | Provides reliable apo-structure models for targets without experimental structures. | |
| Data Resources | Protein Data Bank (PDB) [6] | Repository for 3D structural data of proteins and nucleic acids. | Primary source of experimental structures for target preparation and benchmarking. |
| PDBbind [6] [27] | Curated database of protein-ligand complexes with binding affinity data. | Provides a core set for training and benchmarking scoring functions and AI models. | |
| ChEMBL [6] | Large-scale database of bioactive molecules with drug-like properties. | Source of ligand structures and bioactivity data for model training and validation. | |
| Specialized Analysis | PyMOL / ChimeraX | Molecular visualization system for analyzing structures, poses, and trajectories. | Indispensable for visual inspection of docking results and MD simulation frames. |
| RDKit [27] | Open-source cheminformatics toolkit. | Used for ligand preparation, descriptor calculation, and file format manipulation. |
The landscape of structure-based approaches is defined by a strategic continuum from speed to accuracy. Static molecular docking remains an indispensable tool for the initial exploration of vast chemical space, efficiently prioritizing candidates for more resource-intensive study [24] [23]. Molecular dynamics simulations provide the necessary biophysical depth to validate these candidates, offering unparalleled insights into stability, dynamics, and the thermodynamics of binding [25] [29].
The emerging paradigm, powerfully demonstrated by AI models like DynamicBind, is the fusion of these concepts: achieving dynamic insights at near-docking speeds [27]. This capability to predict ligand-specific protein conformations directly addresses the historical "static receptor" limitation and is particularly promising for targeting cryptic pockets and highly flexible proteins.
Ultimately, the most effective discovery pipeline is not reliant on a single method but on their intelligent integration. This computational cascade should be further informed by and validated with complementary experimental techniques. NMR-driven SBDD, for instance, can provide atomic-level details on dynamics and weak interactions in solution, informing and refining computational models [28]. Furthermore, the dereplication paradigm serves as a crucial checkpoint to ensure that novel structural predictions are translated into genuinely novel chemical matter, avoiding redundant rediscovery [11] [22]. The future of rational drug discovery lies in this synergistic, multi-faceted approach, leveraging the unique strengths of each tool to navigate the complex journey from target structure to viable drug candidate.
This guide compares two fundamental strategies in natural product (NP) research: taxonomy-focused dereplication, which prioritizes the efficient identification of known compounds to avoid rediscovery, and structure-based approaches, which aim to predict bioactivity and mechanism of action (MoA) to discover novel therapeutic leads [30] [22]. Framed within the broader thesis of cataloging known chemistry versus discovering new biology, this analysis provides researchers with a clear comparison of objectives, experimental protocols, performance, and applications.
Core Philosophical and Methodological Comparison
The divergence between these approaches originates from their primary objectives. Taxonomy-focused dereplication is a defensive, efficiency-driven strategy designed to filter out known compounds early in the discovery pipeline. Its core thesis is that focusing on the known chemical space of a specific taxon (species, genus, family) accelerates research by preventing redundant work [11] [31]. In contrast, structure-based discovery is an offensive, novelty-driven strategy. It uses analytical data to prioritize unknown or novel chemical scaffolds, with the explicit goal of uncovering new bioactivities and MoAs, accepting that this may sometimes lead to compounds with no immediate known biological function [22].
The methodological pathways reflect this philosophical split. Dereplication typically begins with a taxonomically defined biological sample, using tools like the LOTUS database to create a focused library of known compounds from related organisms [11]. Structure-based discovery often starts with broad analytical profiling (e.g., LC-MS) of extracts or engineered systems, using computational tools to flag spectral features that do not match known compounds in universal databases [30] [22].
Quantitative Performance Comparison
The following table summarizes the key performance characteristics and outcomes of the two approaches.
| Performance Metric | Taxonomy-Focused Dereplication | Structure-Based Bioactivity Prediction |
|---|---|---|
| Primary Objective | Avoid redundant isolation and characterization of known compounds [11] [31]. | Discover novel chemical scaffolds and predict their biological activity [30] [22]. |
| Typical Success Rate | High identification rate for known compounds within a well-studied taxon [11]. | Lower hit rate for novel bioactive compounds, but higher scaffold novelty [22]. |
| Key Analytical Tools | ¹³C NMR, LC-MS, taxon-specific spectral databases [11] [31]. | HR-MS/MS, Molecular Networking (e.g., GNPS), in silico docking, QSAR models [30]. |
| Time to Initial Result | Rapid (hours to days) for known compound identification [11]. | Longer (days to weeks) for novel compound prioritization and bioassay [30]. |
| Data Integration | Relies on the "Three Pillars": Taxonomy, Molecular Structure, and Spectroscopy [31]. | Integrates genomics, metabolomics, chemoinformatics, and phenotypic screening data [30]. |
| Main Output | Confirmed identity of a known natural product. | Prioritized list of unknown features for isolation & a predicted bioactivity/MoA hypothesis [22]. |
Detailed Experimental Protocols
Protocol 1: Taxonomy-Focused Dereplication via ¹³C NMR Prediction
This protocol, exemplified by the CNMR_Predict workflow for Brassica rapa, details the creation and use of a taxon-specific database for dereplication [11].
Step 1: Taxon-Specific Compound Library Creation
- Query: Perform a search for a target organism (e.g., Brassica rapa) in the comprehensive LOTUS database (https://lotus.naturalproducts.net/).
- Export: Download all associated chemical structures in a standard format (e.g., SDF V3000).
- Clean & Standardize: Use cheminformatics scripts (e.g., RDKit) to remove duplicates, correct tautomeric forms (e.g., converting iminols to amides), and standardize valency representations to ensure compatibility with prediction software [11].
Step 2: Spectral Data Augmentation
- Prediction: Import the cleaned structure library into spectroscopic prediction software (e.g., ACD/Labs CNMR Predictor).
- Calculation: Generate predicted ¹³C NMR chemical shifts for every compound in the library. ¹³C NMR is favored for its wide spectral dispersion and accurate predictability [11].
- Database Compilation: Merge the structural data, taxonomic origin, and predicted NMR shifts into a searchable, taxon-focused database (e.g., a .NMRUDB file).
Step 3: Experimental Sample Analysis & Matching
- Acquisition: Analyze a crude or fractionated extract of the target organism using ¹³C NMR spectroscopy.
- Search: Query the experimental chemical shift list against the custom-built, taxon-focused database.
- Identification: Match the experimental spectrum to a predicted spectrum in the database to confidently identify the known compound, thereby halting further costly isolation efforts [11].
Protocol 2: Structure-Based Workflow for Novel Bioactivity Prediction
This protocol integrates metabolomics and bioinformatics to prioritize novel compounds and suggest their MoA [30] [22].
Step 1: Untargeted Metabolic Profiling
- Analysis: Subject microbial or plant extracts to high-resolution LC-MS/MS analysis.
- Processing: Convert raw data to identify chromatographic peaks (features) with associated m/z and MS/MS fragmentation patterns.
Step 2: Molecular Networking & Novelty Prioritization
- Networking: Upload MS/MS data to the Global Natural Products Social Molecular Networking (GNPS) platform. This clusters compounds with similar fragmentation spectra, visually grouping related molecules [30].
- Dereplication: Annotate nodes (clusters) by matching spectra against public spectral libraries to flag known compounds.
- Prioritization: Target for isolation clusters that are not linked to known compounds or that originate from high-priority biological sources (e.g., uncultured microbes, engineered strains) [22].
Step 3: Bioactivity Prediction & Testing
- In-silico Prediction: For prioritized unknown features, use computational tools to:
- Predict molecular structure from MS/MS fragments or NMR data.
- Perform in-silico docking to suggest potential protein targets.
- Apply Quantitative Structure-Activity Relationship (QSAR) models to forecast biological activity [30].
- Hypothesis-Driven Assay: Design and execute targeted biological screens (e.g., against a specific enzyme or cellular phenotype) based on the computational predictions to validate bioactivity.
Visualizing the Workflows
The following diagrams illustrate the logical flow and key decision points for each methodological approach.
Taxonomy-Focused Dereplication Workflow
Structure-Based Discovery & Bioactivity Prediction
The Scientist's Toolkit: Essential Research Reagents & Materials
Successful implementation of either strategy requires specific tools and resources. The table below details essential solutions for each approach.
| Item | Function in Taxonomy-Focused Dereplication | Function in Structure-Based/Bioactivity Prediction |
|---|---|---|
| LOTUS Database | Primary source for retrieving NP structures linked to a specific taxonomic lineage [11]. | Used for background dereplication within molecular networking to filter known compounds [30]. |
| ACD/Labs CNMR Predictor | Software for generating predicted ¹³C NMR chemical shifts to populate taxon-specific databases [11]. | Less central; may be used later in the pipeline for structural verification of isolated novel compounds. |
| GNPS (Global Natural Products Social) Platform | Can be used to cross-check MS/MS data, but secondary to NMR-based methods. | Core platform for MS/MS data analysis, molecular networking, and community-wide dereplication [30]. |
| RDKit Cheminformatics Toolkit | Used for scripting the cleanup, standardization, and format conversion of structure libraries [11]. | Used for chemical structure manipulation, fingerprint generation, and supporting QSAR/modeling efforts. |
| KnapsackSearch / CNMR_Predict Scripts | Tools for automating the generation of taxon-focused, NMR-augmented databases [11] [31]. | Not typically used. |
| High-Resolution Mass Spectrometer (HR-MS/MS) | Supports identification via exact mass and formula. | Essential for untargeted profiling, generating MS/MS data for networking, and determining molecular formulas [22]. |
| Bioassay Kits & Reagents | Used for general activity screening, but not the primary driver. | Core component. Includes enzyme substrates, cell lines, fluorescent dyes, and reporter systems for HTS and MoA studies [30]. |
| In-silico Docking Software (e.g., AutoDock) | Rarely used. | Predicts the interaction between a putative novel compound and a protein target to hypothesize MoA [30]. |
The choice between taxonomy-focused dereplication and structure-based bioactivity prediction is not mutually exclusive but rather strategic. Taxonomy-focused dereplication excels in efficiency, systematically mapping the chemistry of taxonomic groups and conserving resources [11] [31]. Structure-based approaches excel in novelty discovery, leveraging modern analytics and computation to venture into unknown chemical space with a guided hypothesis for bioactivity [30] [22].
The most effective modern NP discovery programs integrate both philosophies into a single pipeline. Initial rapid dereplication against focused and global databases removes known compounds, while subsequent molecular networking and bioinformatics prioritize the remaining unknown features for isolation. The isolated novel compounds can then be directed toward targeted bioassays based on in-silico MoA predictions. This synergistic strategy, leveraging the strengths of both primary objectives, maximizes the probability of efficiently discovering truly novel and biologically active natural products.
Workflow Deep Dive: From Molecular Networking to Virtual Screening Pipelines
The systematic discovery of novel natural products (NPs) is fundamentally hindered by the challenge of dereplication—the rapid identification of known compounds to avoid redundant rediscovery. Modern dereplication strategies have crystallized into two complementary paradigms: taxonomic-focused and structure-based approaches [22].
The taxonomic-focused approach is historically rooted. It begins with the biological or ecological selection of source material (e.g., a novel microbial species from a unique environment like mangrove sediments [32]), followed by bioactivity-guided fractionation. Chemical analysis is typically performed late in the pipeline, primarily to confirm the structure of an already-isolated bioactive compound. This method risks rediscovery but is driven by specific biological hypotheses.
In contrast, the structure-based approach inverts this workflow. It employs analytical techniques like liquid chromatography-tandem mass spectrometry (LC-MS/MS) at the very beginning to profile the chemical content of a crude extract [22]. The goal is to prioritize unknown chemical entities for isolation before bioactivity testing. This paradigm is powered by three core technologies:
- LC-MS/MS: Provides the sensitive detection and fragmentation data for compounds.
- Molecular Networking via GNPS: Visualizes and clusters MS/MS data to identify novel chemical families and annotate known ones.
- Genome Mining: Predicts the biosynthetic potential of an organism by identifying Biosynthetic Gene Clusters (BGCs) in its genome.
The integration of these tools creates a powerful, hypothesis-driven toolkit for NP discovery. This guide compares the performance, experimental protocols, and synergistic application of these core technologies within the modern structure-based dereplication framework.
Technology Comparison: Core Tools for Dereplication
LC-MS/MS: The Foundational Analytical Engine
LC-MS/MS serves as the indispensable analytical core, generating the primary data upon which molecular networking and integration depend.
- Principle: Separates compounds chromatographically (LC) and then analyzes them via mass spectrometry. The first mass stage (MS1) provides the intact mass-to-charge ratio (m/z) and intensity. The second stage (MS2) fragments selected ions, producing a characteristic spectrum that serves as a structural fingerprint [33].
- Role in Dereplication: Enables the rapid profiling of complex extracts. High-resolution MS1 allows for precise formula prediction, while MS2 spectra are used for database searching and similarity comparisons in molecular networking [22] [34].
Table: Comparison of Key Detection Technologies in Dereplication
| Detection Method | Key Advantages | Primary Limitations | Typical Role in Pipeline |
|---|---|---|---|
| LC-MS/MS | High sensitivity (ng level), provides molecular formula & structural fingerprints, high-throughput compatible [22]. | Ionization bias, requires spectral libraries for confident identification, destructive analysis [22]. | Frontline analysis. Profiling crude extracts, generating data for GNPS and database dereplication. |
| NMR Spectroscopy | Unmatched structural detail, non-destructive, universal for all compounds [22]. | Low sensitivity (mg-µg required), expensive, low-throughput, requires pure compounds [22]. | Late-stage confirmation. Definitive structural elucidation of purified compounds. |
| UV/Vis Spectroscopy | Inexpensive, non-destructive, easily coupled online [22]. | Provides minimal structural information, requires chromophores [22]. | Supplementary detection. Often used in-line with LC for initial profiling. |
Molecular Networking (GNPS): Visualizing Chemical Relationships
Molecular Networking (MN), particularly via the Global Natural Products Social Molecular Networking (GNPS) platform, is a computational-visual tool for organizing and interpreting MS/MS data [34].
- Principle: It operates on the core hypothesis that structurally similar molecules yield similar MS/MS spectra. GNPS calculates spectral similarity (e.g., modified cosine score) between all MS/MS spectra in a dataset and visualizes them as a network where nodes represent spectra and connecting edges represent significant similarity [34] [33]. This clusters analogs, knowns, and unknowns into molecular families.
- Evolution: Classical MN has evolved into more advanced workflows like Feature-Based Molecular Networking (FBMN), which integrates chromatographic information (retention time, peak area) to distinguish isomers and enable quantitative analysis [34] [35].
- Performance: GNPS can achieve limits of detection for specific compounds in complex matrices as low as 0.1–1 ng/g [33]. It significantly increases annotation rates in untargeted studies and has guided the isolation of over 40 known and novel compounds in single studies [34] [33].
Table: Key Molecular Networking Tools and Their Functions
| Tool Name | Type | Primary Function | Key Advantage |
|---|---|---|---|
| Classical MN | Clustering | Groups MS/MS spectra by similarity [34]. | Visualizes chemical relationships, identifies novel clusters. |
| Feature-Based MN (FBMN) | Enhanced Clustering | Integrates LC-MS1 feature data (RT, abundance) [34] [35]. | Handles isomers, links to quantitative data, reduces redundancy. |
| ION Identity MN (IIMN) | Annotation | Groups different ion forms (adducts, dimers) of the same molecule [34]. | Deconvolutes complex MS1 signals, simplifies networks. |
| Network Annotation Propagation (NAP) | Annotation | Propagates annotations within a network based on structural similarity [34]. | Annotates unknown molecules based on known neighbors. |
Diagram 1: A Generalized Molecular Networking Workflow via GNPS. The process begins with LC-MS/MS analysis of a sample, followed by data conversion and upload to the GNPS platform. Core workflows include molecular network construction and library matching, culminating in an annotated network used to prioritize unknown compounds for isolation.
Genome Mining: Predicting Biosynthetic Potential
Genome mining shifts the discovery focus from the expressed metabolite to the genetic potential encoded in an organism's DNA.
- Principle: It involves sequencing an organism's genome and using bioinformatic tools (e.g., antiSMASH) to scan for Biosynthetic Gene Clusters (BGCs)—groups of genes that encode the enzymes for NP biosynthesis [32] [36].
- Role in Dereplication: Identifies strains with high novelty potential (e.g., many unknown BGCs) and provides a genetic hypothesis for the structures one might find (e.g., non-ribosomal peptides, polyketides) [32]. Advanced tools like antiSMASH now include specialized detection modules; for example, its metallophore prediction algorithm achieves 97% precision and 78% recall [36].
- Taxonomic vs. Genomic Selection: While taxonomy can guide strain selection, genome mining is far more predictive. A study on Streptomyces sp. B1866 showed that despite its novel taxonomy, its genome contained 42 BGCs, over half with low similarity to known clusters, correctly predicting chemical novelty later confirmed by isolation [32].
Table: Genome Mining Tools and Comparative Genomics Approaches
| Tool / Approach | Primary Target | Key Metric | Utility in Dereplication |
|---|---|---|---|
| antiSMASH | BGC Identification | BGC count, novelty, class [32] [36]. | Priority ranking. Identifies strains with high/novel biosynthetic potential. |
| skDER / CiDDER | Genomic Dereplication | Average Nucleotide Identity (ANI), protein cluster saturation [37]. | Strain selection. Reduces redundancy in strain collections for sequencing. |
| Alignment-based (AL) Metagenomics | Taxonomic/Functional Profiling | Relative abundance of known taxa/genes [3]. | Community context. Useful for microbiome studies to profile known functions. |
| De novo (DN) Metagenomics | Novel Genome Assembly | Metagenome-Assembled Genomes (MAGs), novel gene discovery [3]. | Novelty discovery. Uncovers novel BGCs from uncultured organisms. |
Integrated Protocol: A Synergistic Workflow
The true power of the modern toolkit is realized through integration. The following detailed protocol, exemplified by the discovery of streptoxazole A from Streptomyces sp. B1866, outlines this synergistic workflow [32].
Stage 1: Strain Selection & Genomic Prioritization
- Strain Isolation & Taxonomic Assessment: Isolate a strain from a unique biotope (e.g., mangrove sediments). Perform 16S rRNA sequencing for preliminary taxonomic placement. For B1866, 16S showed <97% similarity to known species, suggesting a novel Streptomyces sp [32].
- Whole Genome Sequencing & Mining: Sequence the genome. Use antiSMASH to identify and categorize BGCs. Key Decision Point: Prioritize strains with a high number of BGCs, especially those with low similarity (<70%) to known clusters in databases. Strain B1866 was prioritized because its genome contained 42 BGCs, 21 of which were involved in polyketide (PKS), non-ribosomal peptide (NRPS), or hybrid biosynthesis [32].
Stage 2: Metabolomic Analysis & Network-Driven Dereplication
- Cultivation & Extraction: Culture the strain under appropriate conditions (e.g., fermentation). Extract secondary metabolites using organic solvents (e.g., ethyl acetate).
- LC-MS/MS Data Acquisition: Analyze the crude extract using UPLC-MS/MS in data-dependent acquisition (DDA) mode to collect both MS1 and MS2 spectra [32].
- Molecular Networking & Dereplication:
- Process raw data (conversion to .mzML format) and upload to GNPS.
- Perform FBMN analysis and a spectral library search against public repositories.
- Interpretation: Nodes (metabolites) that match library spectra are annotated as known compounds. Nodes that form clusters but have no library match represent novel or rare compound families and are prioritized for isolation [32] [34].
Stage 3: Targeted Isolation & Structure Elucidation
- Guided Isolation: Use the molecular network as a map. Target the precursor ions (m/z) corresponding to prioritized nodes in the network for purification using preparative chromatography.
- Structure Elucidation: Purity is assessed by LC-MS. The structure of the isolated compound is determined using spectroscopic techniques: HRESIMS for molecular formula, and 1D/2D NMR for full structural assignment. For streptoxazole A, this confirmed a novel benzoxazole structure [32].
- Bioactivity Testing: Test purified compounds for biological activity. Streptoxazole A exhibited anti-inflammatory activity with an IC₅₀ of 38.4 μM against LPS-induced NO production [32].
Diagram 2: The Integrated Discovery Workflow. This synergistic protocol begins with genomic assessment to prioritize a strain, uses metabolomics to visualize its chemical output and pinpoint novelty, and culminates in the isolation and characterization of new compounds.
Table: Key Research Reagents and Computational Tools for Integrated Dereplication
| Item / Resource | Category | Function / Purpose | Example / Note |
|---|---|---|---|
| antiSMASH | Software | Identifies and annotates biosynthetic gene clusters in genomic data [32] [36]. | Central tool for genome mining; now includes specialized detectors (e.g., for metallophores [36]). |
| GNPS Platform | Web Platform | Performs molecular networking, library searches, and collaborative annotation of MS/MS data [38] [34]. | Core infrastructure for structure-based metabolomics. |
| UPLC-HRMS System | Instrumentation | Provides high-resolution MS1 and MS2 data for metabolomic profiling and dereplication. | Systems like Q-Exactive Orbitrap are commonly used [32] [35]. |
| Fermentation Media | Reagent | Supports the growth and secondary metabolite production of microbial strains. | Composition is varied (OSMAC approach) to elicit BGC expression [22]. |
| NMR Solvents | Reagent | Required for the final structural elucidation of purified compounds. | Deuterated solvents (e.g., CDCl₃, DMSO-d₆) are essential for 1D/2D NMR experiments [32]. |
| Public Spectral Libraries | Database | Enables dereplication by matching experimental MS/MS spectra to known compounds. | Libraries within GNPS (e.g., MassBank, ReSpect) are critical for annotation [34]. |
| skDER / CiDDER | Software | Performs genomic dereplication to select a non-redundant set of representative genomes for analysis [37]. | Reduces computational burden and bias in comparative genomics. |
The contemporary toolkit for dereplication—LC-MS/MS, GNPS-based molecular networking, and genome mining—transcends the old dichotomy of taxonomic versus structure-based approaches. It forges a unified, iterative framework where each technology informs and validates the others.
Genome mining provides a genetic hypothesis for an organism's chemical capacity, allowing researchers to prioritize the most promising strains before any cultivation. LC-MS/MS and molecular networking then deliver the chemical reality, offering a rapid snapshot of the actual metabolome, dereplicating knowns, and visually highlighting clusters of unknown metabolites for targeted isolation. This integrated workflow, as demonstrated by the discovery of streptoxazole A [32], efficiently bridges an organism's genetic potential with its chemical expression.
The future of NP discovery lies in deepening this integration through automated annotation tools (e.g., DEREPLICATOR+, SIRIUS [34]), large-scale genomic censuses [36], and the application of machine learning to predict molecular properties directly from MS/MS spectra or BGC sequences. For researchers and drug development professionals, mastering this interconnected toolkit is no longer optional but essential for the efficient and targeted discovery of the next generation of natural product-based therapeutics.
The discovery of novel bioactive compounds, such as antibiotics, relies on two primary strategic paradigms: taxonomic-focused dereplication and structure-based drug design (SBDD). These approaches operate on fundamentally different principles and inform distinct stages of the discovery pipeline [4] [11].
Taxonomic-focused dereplication is a front-end strategy aimed at efficiently identifying known compounds within complex biological extracts to prioritize novelty. It leverages the known chemical repertoire of specific organisms (a taxon) to avoid rediscovery. Modern implementations combine advanced cultivation techniques (e.g., microbial diffusion chambers), high-throughput screening, and spectroscopic profiling using tools like mass spectrometry (MS) and nuclear magnetic resonance (NMR), often organized in specialized databases like LOTUS [4] [11]. For instance, an integrated pipeline using diffusion chambers, bioactivity screening, and MS-based dereplication successfully identified both known and novel antibiotic-producing bacteria from soil samples [4].
In contrast, structure-based drug design is a target-driven, back-end approach. It begins with a defined three-dimensional macromolecular target (e.g., a viral protease or a bacterial enzyme) and employs computational tools to design or discover molecules that selectively modulate its function. The core computational toolkit—homology modeling, molecular docking, and molecular dynamics (MD) simulations—allows researchers to predict target structures, simulate ligand binding, and assess complex stability at atomic resolution [39] [40] [41].
This guide provides a comparative analysis of these three pillars of SBDD, benchmarking their performance against alternative methods and framing their application within the broader research context that also includes taxonomic dereplication.
Diagram: The complementary relationship between taxonomic dereplication and structure-based design in drug discovery.
Homology Modeling: Performance and Protocol
Homology modeling, or comparative modeling, predicts a protein's 3D structure based on its amino acid sequence and the known structure of a related template protein. It remains essential for targets lacking experimental structures, though its landscape has been revolutionized by deep learning.
Comparative Performance of Modeling Algorithms
A 2025 comparative study evaluated four modeling algorithms—Homology Modeling (Modeller), Threading, PEP-FOLD3, and AlphaFold—on a set of short, unstable antimicrobial peptides (AMPs). The study used Ramachandran plot analysis, VADAR structure assessment, and 100 ns MD simulations to determine which algorithm produced the most stable conformations for peptides with different physicochemical properties [42].
Table: Performance of Structure Prediction Algorithms for Short Peptides (≤50 aa) [42]
| Algorithm | Core Approach | Optimal Use Case (Peptide Property) | Key Performance Finding | Compact Structure (Avg. over 10 peptides) | Stable Dynamics (Avg. over 10 peptides) |
|---|---|---|---|---|---|
| AlphaFold | Deep Learning (MSA-based) | Hydrophobic peptides | High accuracy for compact structures; complements Threading. | 90% | 60% |
| PEP-FOLD3 | De Novo Folding | Hydrophilic peptides | Provides both compact and dynamically stable structures. | 80% | 80% |
| Threading | Fold Recognition | Hydrophobic peptides | Effective where good templates exist; complements AlphaFold. | 70% | 70% |
| Homology Modeling (Modeller) | Template-based Modeling | Hydrophilic peptides | Reliable when sequence identity to template is high. | 70% | 70% |
Key Findings: The study concluded that no single algorithm was universally superior. Instead, AlphaFold and Threading performed best for more hydrophobic peptides, while PEP-FOLD3 and Homology Modeling were optimal for more hydrophilic peptides [42]. This highlights the need for an integrated, context-dependent approach.
For larger proteins, deep learning models like AlphaFold are now considered state-of-the-art, having largely solved the single-domain protein folding problem. However, challenges persist in modeling large complexes, flexible regions, and the effects of mutations or bound ligands [43]. Emerging tools like DeepFold-PLM address the computational bottleneck of traditional multiple sequence alignment (MSA) generation in AlphaFold, accelerating MSA construction by 47-fold while maintaining comparable prediction accuracy, which is particularly beneficial for high-throughput applications [44].
Experimental Protocol: Building a Homology Model
The following protocol is derived from a 2025 study that created a validated homology model of the human sigma-2 receptor for ligand design [39].
- Template Identification and Alignment: Identify a suitable template structure (e.g., from the PDB) with high sequence similarity to the target. Use the bovine sigma-2 receptor structure (PDB: 7m93) as a template. Align its sequence with the target human sequence (UniProt: Q5BJF2) using alignment software.
- Model Generation: Generate multiple (e.g., five) 3D models using homology modeling software (e.g., Modeller).
- Model Selection and Validation:
- Energetic Screening: Calculate the potential energy of each model and select the one with the lowest energy.
- Steric Validation: Evaluate the selected model's stereochemical quality using a Ramachandran plot (e.g., via PROCHECK). An acceptable model should have >90% of residues in the most favored and allowed regions. The sigma-2 model had 95.9% in allowed regions [39].
- Overall Quality Check: Use scoring functions like the overall quality factor (e.g., from ERRAT). The sigma-2 model scored 87.3% [39].
- Functional Validation via Docking Correlation: Test the model's predictive power by docking a set of ligands with known experimental binding affinities (pKi). Perform a correlation analysis between the computed docking scores and the experimental pKi values. A significant positive correlation (e.g., R² = 0.744 as in the sigma-2 study) supports the model's utility for virtual screening [39].
Molecular Docking: Benchmarking Tools and Rescoring
Molecular docking predicts the preferred orientation and binding affinity of a small molecule (ligand) within a protein's binding site. Its performance varies significantly based on the software and target.
Performance Benchmark of Docking Tools
A 2025 benchmark study evaluated three popular docking tools—AutoDock Vina, PLANTS, and FRED—against wild-type (WT) and drug-resistant quadruple-mutant (Q) variants of Plasmodium falciparum Dihydrofolate Reductase (PfDHFR). Performance was measured using the DEKOIS 2.0 benchmark set and enrichment factor at 1% (EF1%), which indicates the ability to retrieve true active compounds from a large decoy set early in a virtual screening campaign [41].
Table: Benchmarking of Docking and Machine Learning Re-scoring Performance for PfDHFR [41]
| Target | Docking Tool | EF1% (Docking Only) | Best Re-scoring Combination | EF1% (After Re-scoring) | Key Insight |
|---|---|---|---|---|---|
| Wild-Type (WT) | AutoDock Vina | Worse-than-random | Vina + CNN-Score | Better-than-random | ML re-scoring rescued poor initial performance. |
| Wild-Type (WT) | PLANTS | 22 | PLANTS + CNN-Score | 28 | Combined approach yielded best enrichment for WT. |
| Wild-Type (WT) | FRED | 18 | FRED + RF-Score-VS v2 | 24 | Consistent improvement with ML. |
| Quadruple Mutant (Q) | AutoDock Vina | 15 | Vina + RF-Score-VS v2 | 21 | Re-scoring improved enrichment. |
| Quadruple Mutant (Q) | PLANTS | 20 | PLANTS + RF-Score-VS v2 | 27 | Good performance maintained. |
| Quadruple Mutant (Q) | FRED | 23 | FRED + CNN-Score | 31 | Best overall performance for the resistant variant. |
Key Findings:
- Tool Performance is Target-Dependent: No single tool was best for both WT and mutant PfDHFR. FRED combined with CNN-Score performed best for the resistant Q variant (EF1% = 31), while PLANTS with CNN-Score was best for the WT (EF1% = 28) [41].
- Machine Learning Re-scoring Enhances Performance: Applying pretrained ML scoring functions (CNN-Score and RF-Score-VS v2) consistently improved early enrichment over docking alone. In some cases, like with AutoDock Vina on the WT target, it turned worse-than-random screening into a viable one [41].
- Importance for Drug Resistance: The study underscores the need to benchmark docking protocols specifically against resistant mutant targets, as performance can differ significantly from the wild-type [41].
Experimental Protocol: Virtual Screening Workflow
A protocol for structure-based virtual screening (SBVS), integrating docking and MD, is exemplified by a study seeking novel inhibitors for Aeromonas hydrophila [40].
- Target and Library Preparation:
- Obtain the 3D structure of the target protein (e.g., from PDB or homology modeling).
- Prepare a library of candidate ligands (e.g., 100 natural metabolites). Prepare structures by adding hydrogen atoms, assigning charges (e.g., using Gasteiger charges), and minimizing energy.
- Molecular Docking:
- Define the binding site (often based on a known co-crystallized ligand or functional site).
- Dock all library compounds using chosen software (e.g., AutoDock Vina). Use a high exhaustiveness setting for accuracy.
- Rank compounds based on docking score (binding energy in kcal/mol). Select top candidates for further analysis (e.g., Withaferin A: -9.8 kcal/mol vs. control drug Ciprofloxacin: -7.7 kcal/mol) [40].
- Post-Docking Analysis and Filtering:
- Visually inspect the predicted binding poses and interactions (hydrogen bonds, hydrophobic contacts).
- Filter hits based on drug-likeness rules (e.g., Lipinski's Rule of Five) and in silico toxicity predictions.
Diagram: Integrated structure-based virtual screening and validation workflow.
Molecular Dynamics Simulations: Software and Hardware
MD simulations calculate the time-dependent physical movements of atoms, providing insights into the stability, flexibility, and interaction dynamics of protein-ligand complexes.
Comparison of MD Software and Hardware Requirements
MD simulations are computationally intensive. The choice of software and hardware significantly impacts feasibility and turnaround time.
Table: Comparison of Popular Molecular Dynamics Simulation Software [45]
| Software | Primary MD Engine | Key Features | GPU Acceleration | Typical License Model |
|---|---|---|---|---|
| GROMACS | Yes | High performance, excellent parallelization, free & open source. | Yes | Free Open Source (GPL) |
| AMBER | Yes | Comprehensive force fields, widely used in drug discovery. | Yes | Proprietary (free academic) |
| NAMD | Yes | Designed for parallel scaling on large systems. | Yes | Proprietary (free academic) |
| Desmond | Yes | High performance, integrated with Schrödinger suite. | Yes | Proprietary (commercial) |
| OpenMM | Yes | Highly flexible, scriptable Python API. | Yes | Free Open Source (MIT) |
| CHARMM | Yes | Broad force field, long history in academia. | Yes | Proprietary (commercial) |
Table: Recommended Hardware for Molecular Dynamics Simulations (2024-2025) [46]
| Component | Recommended Specifications | Rationale and Notes |
|---|---|---|
| CPU | AMD Ryzen Threadripper PRO or Intel Xeon Scalable. Balance of high clock speed and core count (e.g., 32-64 cores). | MD benefits from parallel processing. High clock speed improves single-threaded performance in preparation steps. |
| GPU | NVIDIA RTX 4090 (24GB VRAM): Best price-to-performance for most systems. NVIDIA RTX 6000 Ada (48GB VRAM): For the largest, most memory-intensive simulations. | GPUs dramatically accelerate the calculation of particle interactions. VRAM capacity limits the size of the simulatable system. |
| RAM | 128 GB - 1 TB (or more) | Must be sufficient to hold the entire simulation system in memory. Large membrane proteins or complexes require >256 GB. |
| Storage | High-speed NVMe SSDs (multiple terabytes) | Fast I/O is critical for writing trajectory files, which can be hundreds of gigabytes. |
Experimental Protocol: Validating a Complex with MD
MD is used to validate the stability of a protein-ligand complex predicted by docking. The protocol below is based on recent studies [39] [40].
- System Preparation:
- Solvation: Place the protein-ligand complex in a simulation box (e.g., a cubic or dodecahedral box) filled with water molecules (e.g., TIP3P water model).
- Neutralization: Add ions (e.g., Na⁺, Cl⁻) to neutralize the system's charge and mimic physiological salt concentration (e.g., 0.15 M NaCl).
- Energy Minimization and Equilibration:
- Minimization: Run a short energy minimization (e.g., 5,000-10,000 steps) to remove steric clashes.
- Equilibration: Perform two equilibration phases in the NVT (constant particles, volume, temperature) and NPT (constant particles, pressure, temperature) ensembles. Gradually heat the system to the target temperature (e.g., 310 K) and stabilize pressure (e.g., 1 bar) over 100-500 ps.
- Production Simulation:
- Trajectory Analysis:
- Root Mean Square Deviation (RMSD): Measure the stability of the protein backbone and ligand pose. A stable complex will plateau at a low RMSD (e.g., ~0.1-0.3 nm for the ligand). In the A. hydrophila study, the complex with Apigenin had a low, stable ligand RMSD of ~0.3 nm over 100 ns [40].
- Root Mean Square Fluctuation (RMSF): Assess flexibility of protein regions (e.g., loop movements).
- Interaction Analysis: Calculate the persistence of key hydrogen bonds and hydrophobic contacts throughout the simulation.
The Scientist's Toolkit: Essential Research Reagent Solutions
This table details essential computational "reagents" and resources required to implement the structure-based design pipeline.
Table: Key Research Reagent Solutions for Structure-Based Design
| Category | Item/Resource | Function/Benefit | Example or Note |
|---|---|---|---|
| Target Structure | Protein Data Bank (PDB) | Repository of experimentally solved protein structures. | Source for templates or direct targets [39]. |
| Modeling Software | AlphaFold Server / ColabFold | State-of-the-art protein structure prediction. | For targets without templates [43] [44]. |
| Modeling Software | Modeller | Gold-standard tool for template-based homology modeling. | Used in the sigma-2 receptor study [39] [42]. |
| Docking Software | AutoDock Vina, PLANTS, FRED | Perform virtual screening by predicting ligand binding. | Benchmarking shows performance is target-dependent [41]. |
| ML Re-scoring | CNN-Score, RF-Score-VS v2 | Pretrained ML models to improve docking hit enrichment. | Significantly improved EF1% in PfDHFR study [41]. |
| MD Software | GROMACS, AMBER, NAMD | Simulate atomic-level dynamics and stability of complexes. | Choice depends on system, force field, and hardware [45]. |
| MD Force Field | CHARMM36, AMBER ff19SB, OPLS-AA | Mathematical parameters defining atomic interactions. | Critical for simulation accuracy. Must match software. |
| Computational Hardware | NVIDIA RTX 40-Series/Ada GPUs | Accelerate docking and MD calculations by orders of magnitude. | RTX 4090 (24GB) and RTX 6000 Ada (48GB) are top choices [46]. |
| Validation Database | DEKOIS, DUD-E | Benchmark sets of known actives and decoys for method validation. | Used to rigorously test docking protocols [41]. |
| Taxonomic DB | LOTUS (Natural Products) | Connects compound structures to taxonomic origin for dereplication. | Used to build taxon-specific NMR databases [11]. |
The field of drug discovery is increasingly defined by its computational methodologies, which serve as the critical bridge between raw chemical data and actionable biological insight. This guide examines the integrated application of Quantitative Structure-Activity Relationship (QSAR) modeling, ADMET (Absorption, Distribution, Metabolism, Excretion, Toxicity) prediction, and machine learning classifiers within modern cheminformatics platforms. These tools are not employed in isolation; their power is unlocked through sophisticated software platforms that manage, analyze, and predict from chemical data [47] [48].
Framed within a broader research thesis comparing taxonomic-focused dereplication and structure-based approaches, this analysis highlights a fundamental strategic divide. Taxonomic methods, prominent in natural product discovery, prioritize biological origin and spectral similarity for dereplication—quickly identifying known compounds within complex extracts [49] [22]. In contrast, structure-based approaches, central to synthetic library screening, begin with the precise chemical structure to predict function, activity, and viability as a drug candidate [50]. The evolution of cheminformatics platforms is, in many ways, a story of convergence, where tools initially designed for one paradigm are now being integrated to serve hybrid workflows, enabling researchers to leverage the strengths of both philosophical starting points.
Comparative Analysis of Leading Cheminformatics Platforms
The selection of a cheminformatics platform is pivotal, as it dictates the scope, efficiency, and innovation capacity of a discovery pipeline. The following table provides a structured, data-driven comparison of leading platforms, evaluating their core competencies in QSAR, ADMET prediction, and machine learning integration.
Table 1: Comparative Analysis of Cheminformatics Platforms for QSAR and ADMET Prediction
| Platform (Vendor) | Core Strength & Licensing | QSAR & SAR Capabilities | ADMET Prediction Features | ML/AI Integration & Specialized Tools |
|---|---|---|---|---|
| RDKit (Open-Source) | Open-source toolkit (BSD); high flexibility & strong community [47]. | Provides descriptors & fingerprints for custom model building; supports MMPA & scaffold analysis [47]. | Computes foundational descriptors (e.g., logP, TPSA); relies on external models for full ADMET prediction [47]. | Seamless integration with Python ML stacks (scikit-learn, PyTorch); serves as the engine for many custom & commercial pipelines [47] [50]. |
| ADMET Predictor (Simulations Plus) | Commercial software specializing in predictive ADMET & PK [51]. | QSAR modeling focused on linking structure to ADMET endpoints. | Flagship capability: >175 predicted properties; models built on proprietary & public data [51]. | In-house developed AI/ML; specialized for pharmacokinetics and toxicity [51]. |
| MOE (Chemical Computing Group) | Comprehensive commercial suite for molecular modeling & drug design [52]. | Integrated QSAR modeling, molecular docking, and structure-based design workflows [52]. | Includes modules for ADMET prediction and property profiling [52]. | Offers machine learning integration and modular workflows for customizable pipelines [52]. |
| deepmirror (deepmirror AI) | Commercial AI platform for hit-to-lead and lead optimization [52]. | Generative AI engine for de novo molecule design and property prediction [52]. | Predicts key ADME and toxicity properties to guide optimization [52]. | Core AI-driven platform; features foundational models for molecule generation and property prediction [52]. |
| Schrödinger (Schrödinger) | Commercial platform combining physics-based & ML methods [52]. | Advanced QSAR via DeepAutoQSAR; FEP for precise binding affinity prediction [52]. | ADMET predictions integrated within lead optimization workflows [52]. | Combines quantum mechanics, ML, and cloud computing for high-throughput simulation [52]. |
Platform Selection and Strategic Fit: The choice of platform is highly contingent on the research paradigm and stage. For taxonomic dereplication in natural product research, open-source and modular tools like RDKit are invaluable. They can be embedded into custom pipelines for processing mass spectrometry data, calculating descriptors for novel scaffolds, and integrating with public spectral libraries like GNPS [49] [34]. Conversely, for structure-based drug design of synthetic compounds, platforms like Schrödinger or ADMET Predictor offer out-of-the-box, validated precision for predicting binding affinities and human pharmacokinetics, which are critical for de-risking candidates before synthesis [51] [52].
A key trend is the rise of federated learning to overcome data limitations, a challenge common to both paradigms. This approach allows multiple institutions to collaboratively train models (e.g., for ADMET) on distributed datasets without sharing raw data, thereby expanding the chemical space covered by the models and improving their generalizability [53].
Experimental Protocols for Model Development and Validation
Robust QSAR and ADMET models rely on rigorous, standardized protocols. Below is a synthesis of best practices for developing and validating predictive cheminformatics models, drawing from recent benchmarking studies.
Table 2: Key Experimental Protocols for QSAR/ADMET Model Development
| Protocol Stage | Key Actions | Purpose & Rationale |
|---|---|---|
| 1. Data Curation & Cleaning | - Standardize SMILES representation [54].- Remove inorganic salts and extract parent compounds [54].- Resolve duplicates, keeping consistent measurements [54]. | Ensures a high-quality, consistent dataset. This step is critical as public datasets often contain inconsistencies that introduce noise and degrade model performance [54]. |
| 2. Molecular Representation (Feature Selection) | - Calculate classical descriptors (e.g., RDKit, topological) [47] [54].- Generate fingerprints (e.g., Morgan/ECFP) [47] [50].- Consider deep learning embeddings (e.g., from GNNs) [50]. | Transforms chemical structures into numerical vectors. A structured approach to testing and combining different representations is needed, as optimal features are often task-dependent [54]. |
| 3. Model Training & Architecture Selection | - Test diverse algorithms (e.g., Random Forest, SVM, Gradient Boosting, GNNs) [50] [54].- Employ hyperparameter optimization (e.g., grid search, Bayesian) [50]. | Identifies the algorithm best suited to capture the structure-activity relationship for the specific endpoint. Ensemble methods often provide robust performance [54]. |
| 4. Validation & Statistical Evaluation | - Use scaffold-based splitting to assess generalizability [54].- Apply cross-validation with statistical hypothesis testing (e.g., paired t-tests) [54].- Evaluate on a hold-out test set and external datasets [50] [54]. | Provides a true estimate of model performance on novel chemotypes. Statistical testing on CV results is more reliable than a single hold-out test score [54]. |
| 5. Practical Performance Assessment | - Evaluate model trained on one data source (e.g., public data) on a different source (e.g., internal assay data) [54].- Use applicability domain analysis to gauge prediction reliability. | Tests model utility in a real-world scenario, where chemical space and assay conditions may differ from training data. This highlights the value of diverse training data via federated learning [53]. |
Visualizing Workflows and Logical Frameworks
From Taxonomic to Structure-Based Dereplication
This diagram illustrates the integrated cheminformatics workflow, highlighting the convergence point where taxonomic and structure-based discovery paradigms meet.
Model Training, Validation, and Application Workflow
This diagram details the systematic process for building, validating, and applying robust machine learning models for property prediction.
The Scientist's Toolkit: Essential Research Reagent Solutions
Beyond software, successful cheminformatics-driven research relies on a suite of essential computational and data resources.
Table 3: Essential Research Reagent Solutions for Integrated Cheminformatics
| Category | Item / Resource | Primary Function & Relevance |
|---|---|---|
| Core Cheminformatics Libraries | RDKit [47], CDK (Chemistry Development Kit) | Open-source toolkits for fundamental operations: molecule I/O, descriptor calculation, fingerprint generation, and substructure search. The foundation for custom pipelines. |
| Specialized Prediction Engines | ADMET Predictor [51], DeepAutoQSAR (Schrödinger) [52] | Provide pre-built, validated models for specific, high-impact endpoints like human pharmacokinetics or toxicity, saving development time. |
| Data Sources & Public Databases | ChEMBL, PubChem, TDC (Therapeutics Data Commons) [54], GNPS [34] | Sources of chemical structures, bioactivity data, and benchmark datasets for model training and validation. GNPS is key for taxonomic/MS-based dereplication. |
| Machine Learning Frameworks | scikit-learn, PyTorch, TensorFlow, Chemprop [50] [54] | Libraries for building, training, and deploying custom ML models. Chemprop is specialized for molecular property prediction using graph neural networks. |
| Workflow & Visualization Tools | KNIME [47], DataWarrior [52], Jupyter Notebooks | Enable the construction of visual, reproducible data pipelines (KNIME), interactive chemical data analysis (DataWarrior), and exploratory coding/analysis (Jupyter). |
| Collaborative Learning Infrastructure | Federated Learning Platforms (e.g., Apheris) [53] | Enables secure, multi-party model training on distributed private datasets, crucial for expanding model applicability domains. |
Thesis Context: Reconciling Taxonomic and Structure-Based Paradigms
The integration of QSAR, ADMET prediction, and machine learning represents a powerful synthesis of the two foundational discovery philosophies. Taxonomic dereplication, exemplified by platforms like GNPS, is inherently data-driven and pattern-based [34]. It uses observed spectral data from complex natural extracts to cluster compounds into molecular families, prioritizing novelty based on spectral dissimilarity to known compounds [49] [22]. Its strength is in efficiently navigating vast biological diversity but can be agnostic to specific biochemical function.
The structure-based approach is fundamentally hypothesis-driven and mechanistic. Starting from a defined molecular structure or protein target, it uses QSAR and physics-based simulations to predict and optimize for a desired function [50] [52]. Its strength is precision and optimization but may overlook serendipitous discoveries from nature's chemical repertoire.
Modern integrated cheminformatics platforms are dissolving this dichotomy. A novel scaffold prioritized via taxonomic dereplication can be instantly profiled using structure-based ADMET predictors to assess its drug-like potential [48]. Conversely, generative AI models trained on structural data can design novel compounds that are then virtually screened for taxonomic novelty against natural product databases. Furthermore, federated learning addresses a universal limitation—scarce and biased data—by allowing models to learn from both proprietary synthetic libraries and diverse natural product screening datasets without compromising privacy [53]. This convergence enables a more holistic discovery strategy, leveraging nature's inspiration while applying rigorous computational filters to de-risk development, ultimately accelerating the journey from hypothesis to viable therapeutic candidate.
The discovery of novel microbiota and their bioactive metabolites from complex environments is a cornerstone of modern microbiology and drug discovery. A central challenge in this field is efficient dereplication—the process of rapidly identifying known organisms or compounds to prioritize truly novel discoveries for further investment. Current research is framed by a pivotal methodological dichotomy: taxonomy-focused approaches versus structure-based approaches [31].
Taxonomy-focused dereplication, exemplified by Metagenome-Assembled Genome (MAG) analysis, prioritizes the genetic identity of microbial lineages. It leverages genomic blueprinting from metagenomic data to uncover uncultured candidate species and their phylogenetic novelty [55] [56]. In contrast, structure-based dereplication focuses on the chemical output of microbes. It uses analytical techniques, primarily mass spectrometry (MS) and nuclear magnetic resonance (NMR) spectroscopy, to identify known metabolites within complex extracts, thereby avoiding the re-isolation of known compounds [11] [8] [31].
This guide provides a comparative analysis of these paradigms. It demonstrates how MAG-based dereplication serves as a powerful, culture-independent tool for expanding the known tree of life and discovering novel microbial taxa. The comparison underscores that while MAGs excel at revealing genomic dark matter, integrated, multi-omic pipelines that combine taxonomic discovery with structural analysis are driving the next frontier in natural product research [4] [57].
Performance Comparison: MAG-based vs. Alternative Dereplication Approaches
The choice of dereplication strategy significantly impacts the efficiency, output, and downstream applicability of microbiome discovery efforts. The table below provides a high-level comparison of the core methodologies.
Table 1: Comparative Overview of Dereplication Approaches for Novel Microbiota Discovery
| Aspect | MAG-based (Taxonomy-Focused) Approach | MS/NMR-based (Structure-Focused) Approach | Integrated Multi-omic/Hybrid Approach |
|---|---|---|---|
| Primary Target | Microbial genomes and taxonomic identity [55] [56] | Chemical structures of microbial metabolites [8] [31] | Both microbial genomes and their metabolic output [4] [57] |
| Key Technology | Shotgun metagenomic sequencing, genome assembly & binning [55] [58] | Mass Spectrometry (MS), Nuclear Magnetic Resonance (NMR) [11] [8] | Combines sequencing, cultivation, MS, and genomics [4] |
| Main Output | High-quality Metagenome-Assembled Genomes (MAGs), phylogenetic novelty [59] | Identified known compounds, annotated spectral networks [8] | Novel isolates, confirmed bioactive compounds, linked BGCs [4] |
| Strength | Culture-independent; reveals vast uncultured diversity ("microbial dark matter") [55] [56] | High-throughput; directly identifies known bioactive chemistries; avoids rediscovery [8] [31] | Links taxonomy to function; validates production; discovers "cryptic" metabolites [4] |
| Limitation | Does not confirm live organism or metabolite production; requires deep sequencing [58] [56] | Misses novel compounds without spectral matches; requires dereplication databases [8] [31] | Technically complex and resource-intensive [4] |
| Typical Application | Expanding microbial tree of life, ecological surveys, genomic potential assessment [55] [59] | Natural product dereplication in drug discovery pipelines [8] [31] | Targeted discovery of novel bioactive agents from specific environments [4] |
The performance of MAG-based dereplication can be further quantified by key experimental outcomes from recent studies, as summarized below.
Table 2: Quantitative Performance Metrics from Key MAG-based and Integrated Studies
| Study Focus | Methodology | Key Quantitative Output | Implication for Dereplication |
|---|---|---|---|
| Human Gut Microbiota Expansion [55] | Assembly of 92,143 MAGs from 11,850 gut metagenomes. | Discovered 1,952 uncultured candidate bacterial species; increased phylogenetic diversity by 281%. | Showcases power of large-scale MAG analysis to dereplicate and expand known taxonomic space. |
| Sequencing Tech Comparison [58] | Parallel MAG recovery from seawater using Illumina (short-read) and PacBio (long-read). | Long-read MAGs had fewer contigs, higher N50, and 88% contained a 16S rRNA gene vs. 23% for short-read. | Long-reads produce higher quality, less fragmented MAGs, improving taxonomic classification. |
| Database Resource (MAGdb) [59] | Curation of a public MAG database. | Contains 99,672 high-quality MAGs (completion >90%, contamination <5%) from 13,702 samples. | Provides a critical resource for dereplication against known genomic diversity. |
| Integrated Antibiotic Discovery [4] | Diffusion chamber cultivation + MS dereplication + genomics. | Recovered 1,218 bacterial isolates; 16% showed antibiotic activity; 33% of bioactive strains dereplicated via MS. | Integrates taxonomy (isolates) and structure (MS) for efficient prioritization. |
| Algorithmic MS Dereplication [8] | DEREPLICATOR+ search of GNPS mass spectra database. | Identified 488 compounds (1% FDR) in Actinomyces spectra, a >5x increase over prior tools. | Highlights efficiency of advanced computational tools for structure-based dereplication. |
Experimental Protocols for Key Methodologies
Core Protocol for MAG-based Dereplication from Complex Samples
The following workflow details the standard process for reconstructing and dereplicating microbial genomes directly from environmental metagenomes [55] [58] [56].
- Sample Collection & DNA Extraction: Collect environmental material (e.g., soil, feces, water). Extract high-molecular-weight genomic DNA. For the human gut study, 13,133 samples were processed [55].
- Shotgun Metagenomic Sequencing: Sequence the DNA using short-read (Illumina) and/or long-read (PacBio, Oxford Nanopore) platforms. Quality trim reads (e.g., using fastp or BBduk) [58].
- De Novo Metagenomic Assembly: Assemble quality-filtered reads into contigs. Common assemblers include SPAdes (for short-reads or hybrids) and Flye (for long-reads) [55] [58]. For example, SPAdes was used with k-mer sizes 21,33,55,77,99,127 for gut metagenomes [55].
- Binning (MAG Reconstruction): Group contigs into putative genomes (bins) based on sequence composition (e.g., k-mer frequency) and abundance across samples. Tools include MetaBAT2, CONCOCT, and MaxBin2. Using multiple binners and integrating results with DAS_Tool improves yields [55] [58].
- Quality Assessment & Filtering: Evaluate bins using CheckM2 or similar to estimate completeness and contamination. Retain only medium-quality (≥50% complete, <10% contaminated) and high-quality (≥90% complete, <5% contaminated) MAGs for downstream analysis [55] [59].
- Dereplication & Taxonomic Classification: Cluster MAGs at a species-level threshold (commonly ≥95% Average Nucleotide Identity over ≥60% of the genome) using tools like dRep to generate a non-redundant set [55] [58]. Classify MAGs taxonomically with GTDB-Tk against the Genome Taxonomy Database (GTDB) [58] [59].
- Novelty Assessment & Downstream Analysis: Identify MAGs with low ANI to reference databases as novel. Analyze their functional potential (e.g., biosynthetic gene clusters with antiSMASH), phylogenetic placement, and ecological prevalence [55] [57].
Protocol for Comparative MAG Recovery Using Different Sequencing Technologies
This protocol from a time-series marine study directly compares MAG quality from short-read (SR) and long-read (LR) data [58].
- Parallel Sequencing: Split extracted DNA from the same sample for both Illumina HiSeq (SR) and PacBio Sequel II HiFi (LR) sequencing.
- Independent Assembly & Binning:
- Quality Control: Refine bins in anvi’o and filter using CheckM criteria (quality score = completeness - 5×contamination ≥50) [58].
- Pairwise Genome Comparison: Identify MAGs recovered by both technologies by calculating Average Nucleotide Identity (ANI) using fastANI. Define "shared" MAGs as those with ≥99% ANI [58].
- Metric Comparison: For shared MAG pairs, statistically compare key metrics: number of contigs, N50 value, genome size, and presence of 16S rRNA gene (detected by barrnap) [58].
Protocol for Integrated Discovery with MS-based Dereplication
This protocol from a soil antibiotic discovery study combines cultivation, bioactivity screening, and structural dereplication [4].
- In Situ Cultivation via Diffusion Chambers:
- Prepare chambers with semi-permeable membranes (0.03 µm). Inoculate with a diluted soil slurry in low-nutrient SMS agar.
- Bury chambers in native soil for 2-4 weeks to allow nutrient and signal exchange.
- Retrieve chambers, extract agar plugs, and domesticate growing colonies on R2A agar to obtain pure isolates [4].
- High-Throughput Bioactivity Screening:
- Culture isolates in 96-well format.
- Use overlay assays with indicator strains (e.g., Staphylococcus aureus, Escherichia coli, including drug-resistant variants) to detect antibiotic production [4].
- MS-based Dereplication of Bioactive Strains:
- Extract metabolites from bioactive culture broth with organic solvents.
- Analyze by Liquid Chromatography-Tandem Mass Spectrometry (LC-MS/MS).
- Process spectra through the Global Natural Products Social Molecular Networking (GNPS) platform. Use workflows like DEREPLICATOR+ to automatically compare spectra against libraries of known natural products and identify known antibiotics (e.g., actinomycin D, valinomycin) [4] [8].
- Genomic Validation and Novelty Probe:
- Sequence the genomes of promising, non-dereplicated isolates.
- Mine for Biosynthetic Gene Clusters (BGCs) using antiSMASH. Correlate BGCs with MS/MS molecular features to target novel compounds [4].
Workflow and Relationship Visualizations
Diagram 1: MAG-based Dereplication Workflow
Diagram 2: Relationship Between Dereplication Paradigms
Diagram 3: Comparative MAG Recovery from Short & Long Reads
Table 3: Key Reagents, Software, and Database Resources for Dereplication Research
| Item Name | Category | Primary Function in Dereplication | Example Use/Citation |
|---|---|---|---|
| SPAdes | Bioinformatics Software | De novo metagenomic assembler for short-read data. | Used to assemble human gut metagenomes prior to binning [55]. |
| MetaBAT2 | Bioinformatics Software | Algorithm for binning assembled contigs into draft genomes. | One of the primary binning tools used in comparative studies [55] [58]. |
| CheckM/CheckM2 | Bioinformatics Software | Estimates completeness and contamination of prokaryotic genomes. | Critical for quality filtering MAGs (e.g., >90% complete, <5% contaminated) [55] [59]. |
| GTDB-Tk & GTDB | Database & Toolkit | Provides standardized taxonomic classification of MAGs. | Used to classify MAGs in the MAGdb and other studies [58] [59]. |
| dRep | Bioinformatics Software | Dereplicates and clusters microbial genomes based on ANI. | Used to generate non-redundant MAG sets at the species level [58]. |
| GNPS Platform | Online Platform & Database | Facilitates mass spectrometry data sharing, molecular networking, and dereplication. | Used for MS/MS spectral analysis to identify known natural products [4] [8]. |
| DEREPLICATOR+ | Bioinformatics Algorithm | Dereplicates MS/MS spectra against databases of natural products. | Identified hundreds of compounds in bacterial extracts, outperforming earlier tools [8]. |
| Diffusion Chamber | Cultivation Hardware | Enables in situ cultivation of unculturable bacteria via nutrient diffusion. | Recovered 1,218 bacterial isolates from soil, enhancing taxonomic diversity [4]. |
| MAGdb | Curated Database | A comprehensive repository of high-quality MAGs from diverse studies. | Contains 99,672 quality-controlled MAGs for comparative analysis and dereplication [59]. |
| R2A & SMS Agar | Growth Media | Low-nutrient media used to cultivate slow-growing or fastidious environmental bacteria. | Used for domesticating diffusion chamber isolates and primary cultivation [4]. |
The search for novel therapeutics from natural sources is guided by two complementary computational philosophies: taxonomy-focused dereplication and structure-based virtual screening.
Taxonomy-focused dereplication is a knowledge-driven approach that prioritizes the rapid identification of known compounds to avoid redundant research. It operates on the principle that taxonomically related organisms produce similar specialized metabolites [11]. This method relies on curated databases linking natural product (NP) structures to their biological sources, such as the LOTUS database [11]. The primary tool is spectroscopic data matching—especially 13C NMR and mass spectrometry—where experimental data from a new extract is compared against predicted or recorded spectra for compounds from related taxa [11] [22]. Its strength is efficiency, but its major limitation is that it is inherently biased towards the rediscovery of known chemical scaffolds.
In contrast, structure-based virtual screening is a hypothesis-driven approach designed to discover novel bioactive compounds. It begins with the three-dimensional structure of a therapeutic target (e.g., a cancer-related protein) and computationally screens vast libraries of small molecules to predict binding [60]. This paradigm does not rely on prior taxonomic knowledge of the compound's source but on the physical-chemical principles of molecular recognition. Advanced implementations use integrated workflows combining pharmacophore modeling, molecular docking, and dynamic simulations to identify promising hits from massive digital libraries, such as the COCONUT database containing over 400,000 natural compounds [61] [62].
This case study focuses on the structure-based paradigm, presenting a comparative guide of contemporary integrated virtual screening studies that have successfully identified natural inhibitors against high-value cancer targets: BCL-2, tubulin, and CDK6.
Comparative Analysis of Virtual Screening Campaigns
The following table summarizes three recent, successful virtual screening campaigns that employed integrated computational strategies to identify natural product inhibitors for distinct cancer targets. The data highlights variations in scale, methodology, and outcomes.
Table 1: Comparative Summary of Integrated Virtual Screening Studies for Cancer Targets
| Screening Aspect | Study 1: BCL-2 Inhibitors [61] | Study 2: Tubulin Inhibitors [63] | Study 3: CDK6 Inhibitors [62] |
|---|---|---|---|
| Primary Target | B-cell lymphoma 2 (BCL-2) | Tubulin (Colchicine site) | Cyclin-dependent kinase 6 (CDK6) |
| Compound Library | COCONUT (407,270 compounds) [61] | Specs Library (200,340 synthetic compounds) [63] | COCONUT (Natural products) [62] |
| Initial Filter | Lipinski’s Rule of Five (276,409 compounds) | N/A | Pharmacophore-based screening (800 compounds) |
| Core Virtual Screening Method | Pharmacophore modeling + Glide (HTVS → SP → XP) docking | Glide molecular docking | Glide molecular docking |
| Key Hit Compounds | CNP0237679, CNP0420384 | Compound 89 (Nicotinic acid derivative) | Four top-ranked compounds (A, B, C, D) |
| Computational Validation | MM-GBSA, DFT, Molecular Dynamics (100 ns) | Molecular docking (EBI competition), Pathway analysis | Molecular Dynamics (100 ns), MM-PBSA, Electrostatic Potential Maps |
| Experimental Validation | In silico ADMET, toxicity prediction | In vitro: Antiproliferation, migration, tubulin polymerization. In vivo: Mouse models, patient-derived organoids. | Binding free energy comparison with standard drug (Palbociclib). |
| Reported Binding Affinity (kcal/mol) | -11.4 to -12.2 (Docking Score) | N/A | Top compound superior to Palbociclib [62] |
| Key Pathway/Effect | Apoptosis induction via BCL-2 inhibition | G2/M arrest, inhibition of PI3K/Akt pathway | Cell cycle arrest via CDK6/Rb-E2F pathway inhibition |
Detailed Experimental Protocols
The following protocols detail the integrated workflows common to the studies compared above.
Protocol 1: Ligand Library Preparation and Pharmacophore-Based Screening
This protocol covers the initial steps to filter a large compound library into a focused set for docking [61] [62].
- Ligand Retrieval: Download the 3D structural database (e.g., COCONUT). Convert all structures to a consistent format (e.g., SDF).
- Pre-processing with LigPrep: Use software (e.g., Schrödinger's LigPrep) to generate correct protonation states at physiological pH (7.0 ± 2.0), generate possible tautomers, and produce low-energy ring conformations [61].
- Drug-Likeness Filtering: Apply Lipinski's Rule of Five to filter out compounds with poor oral bioavailability potential [61].
- Pharmacophore Model Generation:
- Structure-Based: Extract features from a target-ligand co-crystal structure (e.g., from PDB). Identify critical interaction points (H-bond donors/acceptors, hydrophobic regions, aromatic rings) [64].
- Ligand-Based: Align multiple known active compounds to identify common critical chemical features [64].
- Pharmacophore Screening: Use the validated model to screen the filtered library. Compounds that match the pharmacophore hypothesis within a set tolerance (e.g., RMSD < 1.0 Å) are selected for molecular docking.
Protocol 2: Molecular Docking and Binding Affinity Refinement
This protocol describes the docking of filtered hits and the refinement of binding affinity estimates [61] [63] [62].
- Protein Preparation:
- Obtain the target protein's crystal structure (e.g., from RCSB PDB). Remove water molecules and co-crystallized ligands.
- Add hydrogen atoms, assign partial charges, and optimize hydrogen-bonding networks using software (e.g., Schrödinger's Protein Preparation Wizard).
- Define the receptor grid for docking by centering it on the active site of the reference ligand.
- Hierarchical Molecular Docking:
- High-Throughput Virtual Screening (HTVS): Dock the entire pharmacophore-filtered library for a rapid initial ranking.
- Standard Precision (SP) Docking: Re-dock the top-ranked HTVS hits (e.g., 10-20%) for improved accuracy.
- Extra Precision (XP) Docking: Perform a final, rigorous docking on the top SP hits to eliminate false positives and generate a reliable pose ranking.
- Binding Free Energy Estimation (MM-GBSA/MM-PBSA): For the top 10-20 XP poses, perform Molecular Mechanics with Generalized Born Surface Area (MM-GBSA) calculations. This method provides a more accurate estimate of the binding free energy by considering solvation and entropy contributions, offering a superior ranking metric to docking scores alone [61] [62].
Protocol 3: Validation via Molecular Dynamics and DFT
This protocol outlines advanced simulations to validate the stability and properties of top-ranked complexes [61] [62].
- System Setup for MD Simulation: Place the docked protein-ligand complex in a solvation box (e.g., TIP3P water). Add ions to neutralize the system's charge.
- Energy Minimization and Equilibration: Minimize the system energy to remove steric clashes. Gradually heat the system to 310 K and equilibrate under constant pressure (NPT ensemble) for at least 100-200 ps.
- Production MD Run: Run an unrestrained simulation for a minimum of 100 nanoseconds (ns). Use a time step of 2 femtoseconds, saving frames every 10-100 picoseconds for analysis.
- Trajectory Analysis: Calculate the Root Mean Square Deviation (RMSD) of the protein backbone and ligand to assess complex stability. Analyze the Root Mean Square Fluctuation (RMSF) to see changes in protein flexibility. Compute the number of specific hydrogen bonds and contact interactions maintained during the simulation.
- Density Functional Theory (DFT) Analysis: Perform quantum mechanical calculations on the free ligand. Optimize geometry using a functional (e.g., B3LYP) and basis set (e.g., 6-31G(d,p)). Calculate electronic properties like HOMO-LUMO energy gap (indicating chemical reactivity/softness) and molecular electrostatic potential (MEP) surfaces to visualize charge distribution and potential interaction sites [61].
Workflow and Conceptual Diagrams
Diagram 1: Taxonomy-Focused vs. Structure-Based Discovery Workflows
Diagram 2: Integrated Virtual Screening Protocol
The Scientist's Toolkit: Key Research Reagents & Software
Table 2: Essential Computational Tools and Databases for Integrated Virtual Screening
| Tool/Resource Name | Type | Primary Function in Screening | Key Application in Case Studies |
|---|---|---|---|
| COCONUT Database [61] [62] | Database | Provides a vast, curated collection of natural product structures for virtual screening. | Primary compound library for discovering BCL-2 and CDK6 inhibitors. |
| Schrödinger Suite (LigPrep, Glide, Prime) [61] | Software Suite | Performs ligand preparation, molecular docking (HTVS/SP/XP), and MM-GBSA binding energy calculations. | Core platform for the entire virtual screening workflow in BCL-2 and tubulin studies. |
| AutoDock Vina [60] | Software | A widely used, open-source program for molecular docking and virtual screening. | Serves as a common, accessible alternative to commercial docking software. |
| GROMACS/AMBER | Software | Packages for running Molecular Dynamics (MD) simulations to assess complex stability. | Used to validate the stability of top-ranked protein-ligand complexes over time (e.g., 100 ns). |
| RDKit [11] [65] | Cheminformatics Library | Handles chemical informatics tasks, including file conversion, fingerprinting, and pharmacophore feature detection. | Used in database preprocessing and in pharmacophore-guided deep learning models (PGMG). |
| AlphaFold3 [66] | AI Software | Predicts protein-ligand complex structures when experimental structures are unavailable. | Generates reliable holo (ligand-bound) target conformations for docking, improving screening performance. |
| PDBbind Database | Database | A curated collection of protein-ligand complexes with binding affinity data for method training and testing. | Used as a benchmark dataset to train and validate deep learning docking models like Interformer [67]. |
| Gaussian [61] | Software | Performs quantum chemical calculations, including Density Functional Theory (DFT), to analyze electronic properties. | Used to calculate HOMO-LUMO gaps and electrostatic potentials of hit compounds to assess reactivity. |
Overcoming Pitfalls: Benchmarking Databases, Scoring Functions, and Data Integration
The discovery of novel, bioactive natural products (NPs) is a fundamental objective in drug development. This process is critically bottlenecked by dereplication—the early identification of known compounds to avoid costly re-isolation and re-elucidation [30]. For decades, taxonomy-focused dereplication has been a cornerstone strategy, prioritizing novel chemical space based on the evolutionary lineage of the source organism. This approach operates on the premise that taxonomic novelty can proxy for chemical novelty. However, this premise is challenged by significant limitations: 1) Database Bias, where historically studied taxa dominate reference libraries, leaving microbial "dark matter" underrepresented [68]; 2) Novelty Boundaries, where convergent evolution and horizontal gene transfer can lead to identical metabolites in distantly related organisms, or vastly different ones in close relatives [69]; and 3) Silent Gene Clusters, where a genome may encode numerous biosynthetic pathways (Biosynthetic Gene Clusters, BGCs) that are not expressed under standard laboratory conditions, remaining invisible to taxonomy-guided metabolite profiling [70].
This necessitates a paradigm shift towards structure-based approaches, which prioritize the direct analysis of chemical features and genetic architecture over taxonomic origin. This guide provides an objective comparison of these two foundational strategies, framing the analysis within the broader thesis that effective modern discovery requires a synergistic integration of both perspectives to navigate the outlined limitations.
Performance Comparison: Taxonomic vs. Structure-Based Dereplication
The following table summarizes the core performance characteristics of taxonomy-focused and structure-based dereplication methods, highlighting their respective advantages and vulnerabilities in the context of the stated limitations.
Table 1: Comparative Performance of Dereplication Strategies
| Performance Metric | Taxonomy-Focused Dereplication | Structure-Based Dereplication | Supporting Data & Key Limitations |
|---|---|---|---|
| Efficiency & Throughput | High-speed database queries based on taxonomic ID or simple spectral libraries [11]. | Computationally intensive due to complex spectral prediction or genomic mining [8] [70]. | DEREPLICATOR+ searched ~200M spectra in GNPS [8]. antiSMASH analyzes entire genomes for BGCs [69]. |
| Novelty Detection Rate | Prone to missing novel compounds in well-studied taxa and "known" compounds in novel taxa (Novelty Boundary issue). | High potential for identifying novel scaffolds by detecting unique spectral patterns or "silent" BGCs [70]. | ClusterFinder identified >1,000 uncharacterized BGCs in a prominent family, a finding invisible to class-specific tools [69]. |
| Susceptibility to Database Bias | Highly susceptible. Heavily reliant on the completeness and bias of reference databases (e.g., LOTUS, DNP) [11]. | Moderately susceptible. Relies on spectral or genomic libraries, but algorithms can predict features for unrecorded structures [8]. | A global BGC analysis revealed 40% encode saccharides, a class historically underrepresented in NP databases [69]. |
| Ability to Detect Silent Gene Clusters | None. Only detects expressed metabolites. | High. Genomics (e.g., antiSMASH, PRISM) can catalog all BGCs regardless of expression [70]. | Metagenomics of octocorals revealed distinct, host-associated BGC profiles not discernible from environment [68]. |
| Dereplication Confidence | Can be high for well-documented taxa, but annotations may be incorrect if taxonomy is misleading. | Provides direct evidence (MS/MS fragmentation, genomic context) leading to higher-confidence annotations [30] [8]. | DEREPLICATOR+ uses fragmentation graphs and FDR calculation to validate metabolite-spectrum matches [8]. |
Detailed Experimental Protocols
Protocol 1: Taxonomy-Focused Dereplication via 13C NMR Prediction (CNMR_Predict) This protocol uses the LOTUS database and predictive algorithms to create taxon-specific dereplication libraries [11].
- Taxon Definition & Data Retrieval: Define the taxonomic scope (e.g., genus Brassica). Query the LOTUS database via its web interface using the taxon name to download all associated NP structures in SDF format.
- Chemical Library Curation: Process the SDF file using RDKit-based Python scripts (
uniqInChI.py,tautomer.py). This removes duplicate structures, corrects amide tautomers to their dominant form, and adjusts atomic valence notations for compatibility with prediction software. - Spectral Prediction: Import the curated SDF file into ACD/Labs CNMR Predictor and DB software. Execute batch 13C NMR chemical shift prediction using the software's built-in algorithms. Export the enriched database containing structures paired with predicted chemical shifts.
- Dereplication Query: Compare the experimental 13C NMR spectrum of the purified unknown compound or complex mixture against the taxon-specific predicted library. Identification is achieved by matching experimental shifts within a defined error margin (e.g., ± 1 ppm).
Protocol 2: Structure-Based Dereplication via Tandem MS (DEREPLICATOR+) This protocol details the use of the DEREPLICATOR+ algorithm for high-confidence identification from mass spectrometry data [8].
- Spectral Acquisition: Analyze the NP extract using liquid chromatography coupled to high-resolution tandem mass spectrometry (LC-HRMS/MS). Convert raw data to open formats (.mzML, .mgf).
- Fragmentation Graph Construction: For each candidate molecule from a structural database (e.g., AntiMarin, DNP), DEREPLICATOR+ generates a "fragmentation graph." This graph represents the molecular structure as a set of atoms and bonds. It simulates potential fragmentation pathways, creating a theoretical set of fragment ions and neutral losses.
- Spectral Matching & Scoring: The algorithm compares the experimental MS/MS spectrum to the theoretical fragmentation graph of each candidate. It computes a match score based on the shared peaks and their intensities. To control error, it also constructs "decoy" fragmentation graphs by randomizing parts of the molecular graph.
- Statistical Validation: A false discovery rate (FDR) is calculated by comparing the distribution of scores against target and decoy databases. Only matches passing a user-defined FDR threshold (e.g., 1%) are considered valid identifications. High-confidence hits can be used to seed molecular networks to discover structural variants.
Protocol 3: Genome-Mining for Silent BGCs (Integrated omics) This protocol outlines the genomic-led discovery of silent gene clusters and their potential linkage to metabolites [70].
- Genome Sequencing & Assembly: Isolate high-quality genomic DNA from the microbial strain. Sequence using a long-read platform (PacBio, Oxford Nanopore) for contiguity, optionally polished with short-read (Illumina) data for accuracy. Assemble reads into a complete genome or high-quality draft.
- BGC Prediction & Annotation: Annotate the assembled genome using a specialized BGC mining tool such as antiSMASH. The tool uses profile Hidden Markov Models (pHMMs) to identify core biosynthetic domains (e.g., PKS, NRPS) and delineates the boundaries of the surrounding gene cluster. This provides a map of all "silent" and expressed BGCs.
- Metabolomic Profiling: In parallel, cultivate the strain under various conditions and profile the metabolome using LC-HRMS/MS. Process the data to create a molecular network (e.g., in GNPS) to visualize chemical families.
- Integration & Prioritization: Correlate genomic and metabolomic data. BGCs whose predicted product class matches a molecular family in the network are high-priority targets for "awakening" via heterologous expression or targeted cultivation.
Visualization of Key Workflows
Title: Workflow and Limitations of Taxonomy-Focused Dereplication
Title: Integrated Structure-Based and Genomic Dereplication Workflow
The Scientist's Toolkit: Essential Research Reagents & Platforms
Table 2: Key Tools and Resources for Modern Dereplication
| Item / Solution | Function in Dereplication | Relevance to Limitation |
|---|---|---|
| LOTUS Database | A comprehensive, open-source database linking NP structures to taxonomic provenance of source organisms [11]. | Core resource for taxonomy-focused workflows; its completeness dictates vulnerability to Database Bias. |
| ACD/Labs CNMR Predictor | Software for predicting 13C NMR chemical shifts from molecular structure [11]. | Enables creation of taxon-specific spectral libraries for dereplication, circumventing the need for experimental reference data. |
| GNPS (Global Natural Products Social) Molecular Networking | A web-based platform for sharing, processing, and comparing tandem mass spectrometry data to visualize chemical relationships [30] [8]. | A core structure-based ecosystem; molecular networking identifies novel analogs, addressing Novelty Boundaries. |
| DEREPLICATOR+ | Algorithm for identifying NPs from MS/MS spectra by searching against structural databases, covering diverse classes [8]. | A premier structure-based tool that increases dereplication confidence and scope, reducing rediscovery. |
| antiSMASH | The standard genomic platform for the automated identification and annotation of BGCs in microbial genomes [69] [70]. | Critical for mapping the full biosynthetic potential, including Silent Gene Clusters, and guiding targeted discovery. |
| PacBio/Oxford Nanopore Sequencers | Long-read sequencing platforms that generate contiguous DNA sequences, allowing for complete assembly of large BGCs [70]. | Essential technology for obtaining high-quality genomic data to reliably mine for silent BGCs. |
| Integrated Omics Pipelines (e.g., CO-OCCUR) | Algorithms that use co-occurrence patterns of genes to identify BGCs, complementing HMM-based tools like antiSMASH [70]. | Helps uncover non-canonical or novel BGC architectures that might be missed by standard tools, expanding novelty detection. |
The comparative analysis underscores that neither taxonomic nor structure-based dereplication is sufficient in isolation. Taxonomy-focused methods, while efficient, are inherently constrained by historical bias and the imperfect correlation between phylogeny and chemistry. Structure-based approaches, particularly those integrating genomics and metabolomics, directly target the chemical and genetic features of novelty, offering a powerful solution to the challenges of silent gene clusters and complex novelty boundaries.
The future of efficient NP discovery lies in a convergent strategy. This strategy uses taxonomic guidance for initial prioritization of unexplored organisms, thereby mitigating database bias at the source selection stage. It then employs structure-based genomics and metabolomics as the primary engines for characterization, confident annotation, and the targeted activation of silent biosynthetic potential. This synergistic framework promises to accelerate the reliable discovery of novel therapeutic leads from nature's chemical inventory.
Article Thesis and Context
Within the ongoing methodological debate in drug discovery—between taxonomic, sequence-focused dereplication and atomic-resolution, structure-based approaches—this guide provides a critical comparison. Taxonomic methods, which classify and prioritize compounds based on sequence families and evolutionary relationships (e.g., Pfam), offer speed and scalability for novel target identification [71]. In contrast, structure-based methods like molecular docking aim for mechanistic precision by predicting how a small molecule interacts with a protein's three-dimensional form [72]. The central thesis is that while structure-based drug design (SBDD) is indispensable for rational ligand optimization, its practical efficacy is constrained by three interconnected core challenges: the limited accuracy of scoring functions, the difficulties in modeling protein flexibility, and the complex treatment of solvation effects. The resolution of these challenges is pivotal for SBDD to fully realize its potential and provide a decisive advantage over broader, less precise taxonomic filtering methods [6].
Comparative Performance of Docking Methodologies
The accuracy of structure-based methods is not monolithic but varies significantly across different algorithms and use cases. The following tables synthesize quantitative performance data from benchmark studies, highlighting the trade-offs between classical, AI-based, and hybrid docking approaches.
Table 1: Success Rate (SR) Comparison of Docking Methods on the PoseBusters Test Set (n=428 complexes) [73]
| Method Category | Method Name | Required Input | Success Rate (LRMSD ≤ 2 Å) | Key Strength | Key Limitation |
|---|---|---|---|---|---|
| Classical Docking | AutoDock Vina | Native holo structure, defined pocket | 52% | High pose accuracy with ideal input | Dependent on high-quality experimental structure |
| Classical Docking | Gold | Native holo structure, defined pocket | ~45-50% (inferred) | Robust scoring functions | Computationally expensive; rigid protein typically |
| AI-Based Co-folding | Umol (with pocket info) | Protein sequence, ligand SMILES, pocket | 45% | Predicts full flexible complex; no structure needed | Accuracy lower than top classical methods |
| AI-Based Co-folding | RoseTTAFold All-Atom (RFAA) | Protein sequence, ligand data | 42% (8% without templates) | Integrated protein-ligand modeling | Performance drops severely without template info |
| AI-Based Co-folding | Umol (blind, no pocket) | Protein sequence, ligand SMILES | 18% | Completely blind prediction | Low success rate for precise pose |
| Hybrid (AF2 + Docking) | AlphaFold2 + DiffDock | Protein sequence, ligand SMILES | 21% | Uses state-of-the-art predicted structure | Accuracy limited by AF2's apo-state modeling |
Table 2: Performance Metrics Across Key Challenges in Structure-Based Screening [6] [73]
| Challenge | Benchmark/Criteria | Typical Performance Range | Leading Method/Approach | Implication for Drug Discovery |
|---|---|---|---|---|
| Scoring Function Accuracy | RMSD on CASF "Core Set" (~300 complexes) | Pearson R: 0.3 - 0.6 for ΔG prediction | Knowledge-based & ML-scoring functions | Poor affinity ranking leads to false positives/negatives in virtual screening |
| Protein Flexibility (Apo vs. Holo) | Success Rate on apo (unbound) protein structures | Often 50% lower than on holo structures [73] | Ensemble docking, AI co-folding (Umol) | Missed opportunities for novel chemotypes requiring induced fit |
| Solvation & Entropy | Enthalpy-Entropy Compensation | Major source of error in ΔG calculation [72] | Explicit solvent MD, WaterMap | Difficult to optimize for binding selectivity and specificity |
| Generalizability | Performance on unseen target folds | Significant drop vs. trained folds [6] | Physics-based functions generalize better | Limits application to novel target classes (e.g., undrugged families) |
Detailed Experimental Protocols
The comparative data presented above is derived from standardized community benchmarks. Below are the detailed methodologies for two critical types of experiments: evaluating docking pose prediction and assessing binding affinity estimation.
Protocol 1: Evaluation of Docking Pose Prediction (PoseBusters Benchmark)
This protocol outlines the steps for benchmarking docking and co-folding methods on their ability to reproduce experimentally observed ligand poses [73].
Dataset Curation: Compile a non-redundant test set of high-quality protein-ligand complexes from the PDB (e.g., PoseBusters set of 428 complexes). Criteria include:
- Resolution ≤ 2.5 Å.
- Removal of redundant sequences (< 30% identity).
- Validation of ligand chemical correctness and binding site integrity.
Input Preparation:
- For classical docking methods: Prepare the native holo protein structure by removing the ligand, adding hydrogens, and assigning partial charges. The ligand is prepared in its standard 3D conformation.
- For AI co-folding methods: Provide the protein amino acid sequence and the ligand SMILES string. For pocket-informed versions (e.g., Umol-pocket), specify residue indices defining the binding site.
Pose Generation:
- Run the docking or co-folding algorithm using default or recommended parameters.
- For docking, typically generate 10-20 output poses per ligand.
- For co-folding AI, the model outputs a single predicted complex structure.
Pose Assessment:
- Align the predicted protein structure to the experimental reference structure via the protein backbone atoms of the binding site.
- Calculate the Ligand Root-Mean-Square Deviation (LRMSD) between the predicted and experimental ligand heavy atom positions.
- A prediction is deemed a "success" if the LRMSD is ≤ 2.0 Å.
Analysis:
- Calculate the overall Success Rate (SR) as (Number of Successes) / (Total Complexes) * 100%.
- Analyze SR as a function of ligand properties (e.g., flexibility, size) and protein class.
Protocol 2: Evaluation of Scoring Function Affinity Prediction (CASF Benchmark)
This protocol describes the standard method for testing a scoring function's ability to predict experimental binding affinities, a greater challenge than pose prediction [6].
Dataset Curation: Use the PDBbind "Core Set" (e.g., ~300 complexes) curated for the CASF benchmark. This set is designed for minimal interdependence between complexes.
Complex Preparation:
- Use the experimentally determined protein-ligand complex structure.
- Apply a standardized preparation pipeline: add missing hydrogens, optimize side-chain orientations for clashes, and assign consistent atomic charges and force fields.
Scoring:
- Input the prepared complex into the scoring function.
- The function computes a predicted score intended to correlate with the binding free energy (ΔG). No re-docking is performed; the native pose is scored.
Correlation Analysis:
- Obtain the experimental binding affinity (Kd, Ki, or IC50 converted to ΔG) from the PDBbind database.
- Calculate the linear correlation (Pearson correlation coefficient, R) between the set of predicted scores and the experimental ΔG values.
- Perform ranking power tests: Assess the function's ability to correctly rank the ligands within a series for a single protein target.
Validation:
- Ensure strict separation between the benchmark set and any data used to train the scoring function to avoid overfitting.
- Cross-validate results across different protein families to assess generalizability.
Visualizing Core Concepts and Workflows
The following diagrams, generated using Graphviz DOT language, illustrate the logical relationships and experimental workflows central to the comparison between taxonomic and structure-based approaches, as well as the specific challenges in SBDD.
Diagram 1: 52 chars - Taxonomic vs. Structure-Based Drug Discovery Workflow
Diagram 2: 52 chars - Scoring Function Composition and Error Sources
Diagram 3: 49 chars - The Protein Flexibility Challenge in Docking
Table 3: Key Databases and Software for Structure-Based Methods Research
| Item Name | Type | Primary Function in Research | Key Consideration for Use |
|---|---|---|---|
| Protein Data Bank (PDB) | Database | Primary repository for experimentally determined 3D structures of proteins and complexes [72] [6]. | Quality varies (resolution, completeness); requires preprocessing (adding H+, fixing residues). |
| PDBbind / CASF Benchmark | Curated Database & Benchmark | Provides curated protein-ligand complexes with binding affinity data for training and fair testing of scoring functions [6]. | Essential for method validation; "Core Set" is standard for affinity prediction tests. |
| ChEMBL / BindingDB | Bioactivity Database | Massive repositories of ligand bioactivity data (Ki, IC50, etc.) against targets [6]. | Critical for knowledge-based/AI model training; data from diverse sources requires careful filtering. |
| AlphaFold2 Protein Structure Database | Prediction Database | Provides highly accurate predicted protein structures for targets with no experimental model [73]. | Predictions are often in apo state, which may not be suitable for docking without refinement. |
| AutoDock Vina, GOLD, Glide | Docking Software | Widely used classical docking programs for pose prediction and virtual screening [72] [73]. | Performance depends on input structure quality and parameter tuning. Treat protein as rigid. |
| GROMACS, AMBER, NAMD | Molecular Dynamics (MD) Software | Simulate protein, ligand, and solvent dynamics to study flexibility and binding thermodynamics [6] [74]. | Computationally expensive; used for refinement and detailed mechanistic studies, not primary screening. |
| Umol, RoseTTAFold All-Atom | AI Co-folding Software | Predict the joint 3D structure of a protein-ligand complex directly from sequence and SMILES [73]. | Emerging tool; promising for flexibility but may lag in pose accuracy vs. classical docking with holo structures. |
| RDKit | Cheminformatics Toolkit | Open-source library for handling ligand preprocessing, force field assignment, and chemical descriptor calculation. | De facto standard for ligand preparation and manipulation in computational chemistry workflows. |
The discovery of novel bioactive natural products (NPs) is persistently hindered by the costly and time-consuming process of dereplication—the early identification of known compounds to avoid redundant characterization [30]. This challenge sits at the heart of a fundamental strategic divide in NP research: the choice between taxonomy-focused and structure-based dereplication approaches. Taxonomy-focused methods leverage the evolutionary principle that related organisms produce similar metabolites, using biological origin as a primary filter to narrow candidate lists [11]. In contrast, structure-based approaches prioritize analytical data (e.g., MS/MS fragmentation, NMR shifts) to identify compounds irrespective of source, often enabling the discovery of structurally novel scaffolds with potentially new modes of action [22].
The integration of Advanced Molecular Networking (MN), particularly Feature-Based Molecular Networking (FBMN), with multi-omics data (genomics, metabolomics) represents a paradigm shift, merging the strengths of both philosophies [30] [75]. This guide provides a comparative analysis of modern dereplication workflows, evaluating their performance through experimental data and detailing the protocols that enable researchers to navigate the trade-offs between efficiency and novelty in NP discovery.
Comparative Analysis of Dereplication Workflows
The selection of a dereplication strategy significantly impacts the throughput, novelty rate, and resource allocation of a discovery campaign. The table below quantifies the performance characteristics of three dominant approaches.
Table 1: Performance Comparison of Dereplication Strategies
| Strategy | Core Technology | Novelty Identification Rate | Throughput (Samples/Week) | Key Limitation | Best For |
|---|---|---|---|---|---|
| Taxonomy-Focused NMR [11] | 13C NMR databases curated by taxon (e.g., LOTUS, CNMR_Predict). | Low to Moderate. Highly effective for known taxa; novelty is a negative result. | Moderate (10-100). Limited by NMR acquisition time and database scope. | Requires high-quality, concentrated isolate; limited to compounds in taxonomic DB. | Rapid confirmation of known compounds in well-studied biological families. |
| Traditional MS Dereplication [22] | LC-HRMS (MS1) for exact mass & formula matching against DBs. | Low. Prone to missing isomers and novel compounds with known formulas. | High (100+). Automated LC-MS runs with fast DB queries. | Depends on mass accuracy; cannot distinguish isomers; provides minimal structural insight. | High-volume pre-screening to filter out obvious knowns from large extract libraries. |
| Advanced MN & Multi-Omics [30] [75] | LC-MS/MS (FBMN) integrated with genomic data. | High. Networks visualize novelty as disconnected clusters; genomics predicts new scaffolds. | Moderate to High (50-200). Computational bottleneck is networking analysis. | Complex data analysis; requires bioinformatics expertise; analog annotations can be tentative. | Discovery-driven projects aiming for new chemical scaffolds and biosynthetic gene cluster (BGC) linkage. |
Experimental Data Supporting Comparison: A landmark study underpinning FBMN reported that molecular networking of 196 fungal extracts led to the annotation of 76% of MS/MS spectra and, crucially, the targeted isolation of a previously overlooked novel antifungal compound, guided by a distinct cluster in the network [75]. In contrast, a taxonomy-focused 13C NMR study on Brassica rapa efficiently dereplicated 121 known compounds but was not designed for de novo discovery [11]. Crucially, the integration with genomics, such as linking a molecular family to a cryptic biosynthetic gene cluster via pattern correlation, can increase the confidence in targeting truly novel metabolites for isolation [30] [22].
Experimental Protocols for Integrated Workflows
Protocol: Feature-Based Molecular Networking (FBMN) on GNPS
This protocol is adapted from the Global Natural Products Social Molecular Networking (GNPS) platform guidelines [75].
1. Sample Preparation & LC-MS/MS Acquisition:
- Prepare crude extracts or fractionated samples. Use reversed-phase LC (e.g., C18 column) coupled to a high-resolution tandem mass spectrometer (e.g., Q-TOF, Orbitrap).
- Acquire data in data-dependent acquisition (DDA) mode. Collect full MS scans (e.g., m/z 100-1500) and top N MS/MS scans per cycle.
2. Data Processing with MZmine or MS-DIAL:
- Process raw data (.mzML files) using MZmine, MS-DIAL, or other supported tools.
- Key steps: Chromatogram building, feature detection (identification of m/z-retention time pairs), deisotoping, alignment across samples, and gap filling.
- Export two files: (A) A feature quantification table (.CSV) listing features with m/z, RT, and intensity per sample. (B) An MS/MS spectral summary file (.MGF) containing fragmentation spectra for each feature.
3. Molecular Networking Job on GNPS:
- Access the FBMN workflow on the GNPS website (requires login).
- Upload the feature table and .MGF file.
- Set critical parameters:
- Precursor Ion Mass Tolerance: 0.02 Da (for high-res instruments).
- Fragment Ion Mass Tolerance: 0.02 Da.
- Min Pairs Cosine: 0.7 (controls edge creation threshold).
- Minimum Matched Peaks: 6.
- Library Search: Enable to annotate nodes against spectral libraries.
- Submit the job. Results include an interactive molecular network and annotated feature tables.
Protocol: Integrating Genomics via BGC-Metabolite Correlation
1. Genome Sequencing and BGC Prediction:
- Sequence the source organism’s genome (e.g., Illumina/PacBio).
- Use antiSMASH or similar software to identify and annotate Biosynthetic Gene Clusters (BGCs).
2. Metabolite Feature Prioritization from FBMN:
- From the FBMN results, identify molecular families (clusters) with no or poor library matches, indicating potential novelty.
- Within these families, prioritize features that show correlation with specific cultivation conditions or that are unique to a single producing strain.
3. In-silico Cross-Domain Analysis:
- Compare the retention time and MS/MS patterns of the prioritized metabolite family with the predicted chemical class (e.g., non-ribosomal peptide, polyketide) from a BGC of interest.
- Use tools like BiG-SCAPE to compare the BGC against databases; unique or hybrid BGCs strengthen the case for novelty.
- This correlation creates a testable hypothesis: that the target BGC produces the metabolites in the target molecular family.
Visualizing the Integrated Dereplication Pipeline
The following diagrams, created using Graphviz DOT language, map the logical workflow of an integrated dereplication strategy and the decision framework for choosing between taxonomic and structure-based approaches.
Diagram 1: Integrated Multi-Omics Dereplication Workflow (Max width: 760px)
Diagram 2: Dereplication Strategy Decision Framework (Max width: 760px)
The Scientist's Toolkit: Essential Reagents & Materials
Table 2: Key Research Reagent Solutions for Advanced Dereplication
| Item | Function in Dereplication | Example / Specification |
|---|---|---|
| LOTUS Database [11] | A comprehensive, open-access database linking NP structures to taxonomic data. Enables taxonomy-focused searches. | lotus.naturalproducts.net |
| ACD/Labs CNMR Predictor [11] | Software for predicting 13C NMR chemical shifts. Used to supplement taxon-specific databases when experimental reference data is missing. | Commercial software for structure-based shift prediction. |
| GNPS Platform [30] [75] | The central, cloud-based ecosystem for performing FBMN, library searches, and sharing spectral data. | gnps.ucsd.edu |
| MZmine / MS-DIAL [75] | Open-source software for processing LC-MS/MS data into feature tables and MS/MS spectral files suitable for GNPS. | Critical for data preparation before molecular networking. |
| antiSMASH [30] | The standard bioinformatics tool for genome mining and the identification of Biosynthetic Gene Clusters (BGCs). | Enables the genomics arm of multi-omics integration. |
| Reversed-Phase LC Column | Separates complex natural extracts prior to MS analysis. Column chemistry drastically affects metabolite coverage. | e.g., C18 column, 2.1 x 100 mm, 1.7 µm particle size. |
| High-Resolution Mass Spectrometer | Provides the accurate mass and fragmentation data (MS/MS) essential for FBMN and formula prediction. | e.g., Q-TOF or Orbitrap-based instruments. |
| Silica & Sephadex LH-20 | Standard chromatography media for the targeted isolation of compounds prioritized from networking or genomic clues. | Used for offline fractionation following bio- or chemi-guided isolation. |
The relentless pursuit of novel therapeutics demands ever more efficient strategies to navigate vast chemical and biological space. In early drug discovery, this challenge manifests as a strategic dichotomy between taxonomy-focused dereplication and structure-based virtual screening. Dereplication strategies, which aim to rapidly identify known compounds from complex mixtures such as natural product extracts, rely heavily on taxonomic and spectroscopic databases (e.g., LOTUS) and predictive tools (e.g., CNMR_Predict) to avoid redundant rediscovery [11]. In contrast, structure-based approaches leverage the three-dimensional architecture of a biological target to rationally select or design novel binders. This comparison guide focuses on the latter paradigm, dissecting the performance of three advanced computational methodologies that are reshaping virtual screening: ensemble docking, alchemical free energy calculations, and AI-enhanced scoring. These techniques address the critical limitations of conventional single-structure docking—primarily its inability to model protein flexibility, accurately rank compounds by affinity, and efficiently scale to ultra-large libraries. Framed within the broader research context that also values taxonomic intelligence, this guide provides an objective, data-driven comparison of these cutting-edge computational tools, equipping researchers to select and integrate optimal strategies for their specific drug discovery campaigns [76] [77] [78].
Performance Comparison of Virtual Screening Methodologies
The following tables provide a quantitative comparison of the performance, typical use cases, and resource requirements for the three core methodologies and their leading implementations.
Table 1: Performance Benchmarks of Virtual Screening Methodologies
| Methodology | Typical Application | Key Performance Metric | Reported Performance | Reference |
|---|---|---|---|---|
| Ensemble Docking (for IDPs) | Hit ID for intrinsically disordered proteins (e.g., α-synuclein) | Correlation with NMR binding affinities | Accurate ranking of high-/low-affinity ligands for α-synuclein fragment [76] | [76] |
| AI-Enhanced Scoring (RosettaVS) | Virtual screening of ultra-large libraries | Top 1% Enrichment Factor (EF1%) on CASF2016 | EF1% = 16.72 (outperformed 2nd best: 11.9) [77] | [77] |
| AI-Enhanced Scoring (RosettaVS) | Pose prediction (docking power) | Success rate (RMSD ≤ 2Å) | Superior performance in binding funnel analysis [77] | [77] |
| Pose Ensemble GNN (DBX2) | Binding affinity prediction | Pearson's R vs. experimental ΔG | R = 0.85 on hold-out test set [79] | [79] |
| Pose Ensemble GNN (DBX2) | Retrospective virtual screening | AUC (Area Under ROC Curve) | AUC = 0.91 on LIT-PCBA subset [79] | [79] |
| Pharmacophore AI (Alpha-Pharm3D) | Bioactivity prediction & screening | Mean AUROC across diverse targets | AUROC ~0.90 [80] | [80] |
| Free Energy Perturbation (FEP+) | Potency & selectivity optimization | Mean Absolute Error (MAE) vs. experiment | ~1.0 kcal/mol (equivalent to 6-8 fold in Ki) [78] | [78] |
| Conventional Docking (Baseline) | General virtual screening | CNN Score Cutoff for quality | CNN score >0.9 improves specificity of candidate selection [81] | [81] |
Table 2: Computational Cost and Typical Use Case Comparison
| Methodology | Computational Cost | Typical Time Scale | Primary Strength | Key Limitation |
|---|---|---|---|---|
| Ensemble Docking | Moderate-High (requires ensemble generation) | Hours to days | Models protein flexibility/heterogeneity; essential for IDPs [76]. | Quality depends on input conformational ensemble [76]. |
| Alchemical Free Energy (FEP+) | Very High | Hours per prediction | High accuracy for relative binding affinity; direct thermodynamic basis [78]. | Requires high-quality structure; complex setup; not for initial screening. |
| AI-Enhanced Scoring | Low (scoring) to Moderate (training) | Seconds to minutes per compound | High speed & excellent ranking for ultra-large libraries; integrates flexibility [77]. | Generalizability to novel scaffolds/ targets can be limited [79]. |
| Pose Ensemble GNN (e.g., DBX2) | Moderate (requires pose generation) | Minutes per compound | Learns from multiple poses; strong affinity prediction [79]. | Dependent on initial docking poses; requires curated training data. |
| Pharmacophore AI (e.g., Alpha-Pharm3D) | Low-Moderate | Seconds per compound | Interpretable 3D pharmacophore models; scaffold hopping [80]. | Performance depends on quality and diversity of training ligands [80]. |
Detailed Experimental Protocols
To ensure reproducibility and provide clear insight into how key performance data was generated, this section outlines the experimental protocols from seminal studies for each methodology.
- Objective: To predict binding modes and relative affinities of small molecules to the intrinsically disordered C-terminal fragment of α-synuclein (residues 121-140).
- Protein Ensemble Preparation:
- Input conformational ensembles were derived from long-timescale all-atom Molecular Dynamics (MD) simulations (100-200 μs) performed with the a99SB-disp force field and validated against NMR data (chemical shifts, scalar couplings, RDCs).
- Snapshots were extracted from simulations of the apo protein and, where available, from ligand-bound simulations.
- Ligand Preparation:
- Test ligands (Fasudil, high-affinity Ligand 47, low-affinity Ligand 23) were prepared in their dominant protonation states at physiological pH.
- Docking Execution:
- Protocol 1 (Force Field-Based): Docking was performed against each protein snapshot in the ensemble using AutoDock Vina. A consensus score (e.g., average or minimum binding energy) across the ensemble was calculated for each ligand.
- Protocol 2 (AI-Based): The same ensemble was docked using DiffDock, a deep learning diffusion model.
- Validation:
- Affinity Ranking: The consensus scores from both protocols were compared to the relative binding affinities measured by NMR spectroscopy.
- Pose Validation: The distribution of docked poses was clustered and compared to the binding modes observed in the long-timescale MD simulation trajectories using t-SNE analysis.
- Objective: To screen multi-billion compound libraries against structured targets (KLHDC2, NaV1.7) with high accuracy and speed.
- Workflow:
- Phase 1 - Fast Prescreening (VSX Mode): A modified, fast protocol of Rosetta GALigandDock was applied to the entire library using a rigid receptor.
- Phase 2 - Active Learning: A target-specific neural network was trained on-the-fly to predict docking scores, iteratively selecting the most promising compounds for full docking calculations.
- Phase 3 - Refined Ranking (VSH Mode): The top hits from Phase 1 were re-docked using the high-precision protocol (VSH) with full receptor flexibility (side-chain and limited backbone movement).
- Scoring: The final ranking used RosettaGenFF-VS, a physics-based scoring function combining enthalpy (ΔH) and a newly developed entropy (ΔS) model.
- Benchmarking: The protocol was validated on the CASF2016 and DUD datasets, measuring docking power (pose prediction) and screening power (enrichment factors).
- Objective: To discover potent and selective Wee1 kinase inhibitors by optimizing for kinome-wide selectivity.
- Workflow:
- Step 1 - Potency Identification (L-RB-FEP+): Billions of design ideas were filtered, and ~9000 were selected for ligand-relative binding Free Energy Perturbation (FEP+) calculations in the Wee1 binding site. Predictions within ~1.0 kcal/mol of experiment were considered reliable.
- Step 2 - Primary Selectivity Check: Compounds predicted to be potent against Wee1 were profiled using L-RB-FEP+ in the binding site of a key off-target (PLK1).
- Step 3 - Kinome-Wide Selectivity Profiling (PRM-FEP+): For promising series, protein residue mutation FEP+ was used. The Wee1 gatekeeper residue (Asn) was alchemically mutated in silico to residues found in other kinases (e.g., Thr, Val, Phe). The calculated change in binding energy (ΔΔG) for a ligand predicted its loss of potency against kinases with that different gatekeeper.
- Step 4 - Experimental Validation: Synthesized compounds were tested in biochemical assays for Wee1 potency and profiled in a panel of 403 wild-type human kinases (DiscoverX scanMAX) to confirm selectivity patterns predicted by PRM-FEP+.
The following diagrams, generated using Graphviz DOT language, illustrate the logical workflows of the core methodologies and their place in the virtual screening ecosystem.
Diagram 1: Method Selection Workflow (94 characters)
Diagram 2: AI & Physics Integration (92 characters)
Table 3: Key Software Tools and Resources for Advanced Virtual Screening
| Category | Tool/Resource Name | Primary Function | Key Feature / Note |
|---|---|---|---|
| Docking & Scoring Engines | AutoDock Vina [76] [81] | Molecular docking and scoring. | Widely used, free, force-field based. Basis for many enhancements. |
| DiffDock [76] | AI-based molecular docking. | Diffusion model for blind pose prediction; used in IDP ensemble studies. | |
| RosettaVS (Rosetta GALigandDock) [77] | High-accuracy docking & scoring. | Models receptor flexibility; high enrichment in benchmarks. | |
| GNINA [81] | Docking with CNN scoring. | Provides CNN score (0-1) for pose quality assessment; improves specificity. | |
| AI & Machine Learning Models | DockBox2 (DBX2) [79] | Graph Neural Network for rescoring. | Uses ensembles of docking poses to predict affinity and pose likelihood. |
| Alpha-Pharm3D [80] | 3D Pharmacophore prediction & screening. | Creates interpretable pharmacophore models from ligand/receptor data. | |
| Free Energy Calculations | FEP+ (Schrödinger) [78] | Relative binding free energy calculations. | Industry standard for high-accuracy ΔΔG prediction (~1 kcal/mol error). |
| Conformational Sampling | Molecular Dynamics (e.g., AMBER, GROMACS) [76] | Generating protein conformational ensembles. | Essential for ensemble docking, especially for flexible/IDP targets. |
| Structure Prediction | AlphaFold2 [82] | Protein structure prediction. | Provides reliable models for targets without experimental structures. |
| Compound Libraries | ZINC, Enamine REAL [77] [81] | Sources of purchasable compounds for screening. | Ultra-large libraries (billions) are now accessible for virtual screening. |
| Experimental Validation | NMR Spectroscopy [76] | Measuring ligand binding to IDPs. | Key validation method for disordered protein interactions. |
| Kinase Profiling Panels (e.g., DiscoverX) [78] | Measuring kinome-wide selectivity. | Essential for experimental validation of off-target predictions (403 kinases). |
The landscape of virtual screening is no longer dominated by a single technique but is a collaborative ecosystem of specialized tools. Ensemble docking is the definitive solution for tackling highly flexible targets, especially intrinsically disordered proteins, where traditional methods fail [76]. For the Herculean task of screening ultra-large chemical libraries, AI-enhanced scoring and active learning platforms like RosettaVS and DBX2 offer an unmatched combination of speed and ranking accuracy, making billion-compound screens feasible in days [77] [79]. When the campaign advances to optimizing a lead series for potency and, crucially, kinome-wide selectivity, alchemical free energy calculations (FEP+) provide the gold standard in predictive accuracy, enabling rational design with reduced experimental cycles [78].
The most powerful strategy is a convergent one. A typical pipeline might begin with AI-accelerated screening of a vast library against an ensemble of target structures to identify diverse hits. These hits could be rescored with pose-ensemble GNNs like DBX2 for improved affinity prediction. Finally, the most promising chemotypes can be optimized using FEP+ calculations to dial in potency and selectivity before synthesis. This integrated approach, which combines the scalability of AI with the rigor of physics-based methods, represents the current frontier in computational drug discovery. By understanding the strengths, costs, and optimal applications of each method detailed in this guide, research teams can make informed decisions that accelerate the journey from target to candidate.
The identification and characterization of biomolecules—whether for understanding microbial diversity or for rational drug design—increasingly relies on computational interrogation of large-scale data repositories. This process hinges on two fundamentally distinct strategies: taxonomic-focused dereplication and structure-based approaches. Taxonomic dereplication reduces redundancy in genomic or proteomic datasets by clustering sequences based on similarity, aiming to select a representative subset that captures the diversity of a sample or population [37]. In contrast, structure-based methods leverage the three-dimensional architecture of proteins to infer function, predict interactions, and guide the design of molecular probes or therapeutics [6].
The efficacy of both paradigms is critically dependent on the quality and comprehensiveness of their underlying reference databases. For dereplication, this means curated, non-redundant sequence libraries [83]. For structural approaches, it requires accurate, high-resolution models of proteins and their complexes [84]. Advances in machine learning and high-throughput experimentation are transforming both fields: deep learning now enables the in silico prediction of mass spectra from peptide sequences to build spectral libraries [85] [86], while AI-driven structure prediction has populated databases with hundreds of millions of protein models [87]. This guide objectively compares the tools, databases, and experimental outcomes associated with these two approaches, providing researchers with a framework to select the optimal strategy for their biological questions.
The choice between dereplication and structure-based analysis is dictated by the research goal, available data, and desired outcome. The following tables summarize the core algorithms, key databases, and performance metrics associated with each approach.
Table 1: Comparison of Core Algorithmic Strategies
| Aspect | Taxonomic-Focused Dereplication | Structure-Based Approaches |
|---|---|---|
| Primary Objective | Reduce genomic/proteomic redundancy; select representative sequences [37]. | Predict function, interactions, and ligands from 3D protein structure [87] [6]. |
| Foundational Data | Nucleotide or amino acid sequences. | Atomic coordinates (experimental or predicted), ligand binding data [6]. |
| Key Metrics | Average Nucleotide Identity (ANI), Alignment Fraction (AF), protein cluster saturation [37]. | Template Modeling Score (TM-score), pLDDT, binding affinity (Kd/Ki), docking score [87] [84]. |
| Typical Workflow | 1. Calculate pairwise similarity (e.g., ANI). 2. Cluster based on thresholds. 3. Select representative(s) [37]. | 1. Acquire/predict structure. 2. Identify functional sites. 3. Dock ligands/virtual screen [88]. |
| Major Tools | skDER, CiDDER, FastANI, CD-HIT [37]. | AlphaFold, RoseTTAFold, molecular docking suites (e.g., AutoDock), FEP tools [6] [84]. |
| Strengths | Fast, scalable for thousands of genomes; clear operational taxonomic units. | Provides mechanistic insight; enables rational design for drug discovery [88]. |
| Limitations | Limited functional insight; dependent on threshold selection. | Computationally intensive; accuracy depends on structure quality [84]. |
Table 2: Key Reference Databases and Data Sources
| Database Name | Primary Content | Applicable Approach | Key Features & Access |
|---|---|---|---|
| UniProt | Comprehensive protein sequences & annotations [89]. | Dereplication, foundation for both. | Expertly curated (Swiss-Prot) and automated (TrEMBL) sections. |
| PRIDE / Peptide Atlas | Mass spectrometry-derived peptide spectra & identifications [89]. | Spectral library building, empirical validation. | Raw data repository; Peptide Atlas uses uniform reprocessing pipeline. |
| AlphaFold DB (AFDB) | AI-predicted protein structure models for millions of sequences [87]. | Structure-based analysis. | Covers vast sequence space; includes confidence metric (pLDDT). |
| ESMAtlas | High-quality predicted structures for metagenomic proteins [87]. | Structure-based analysis, metagenomics. | Focus on prokaryotic and environmental sequences. |
| Protein Data Bank (PDB) | Experimentally determined 3D structures [6] [84]. | Structure-based validation & modeling. | Gold standard for experimental structures; may contain artifacts. |
| ChEMBL / BindingDB | Bioactivity data for drug-like molecules against targets [6]. | Structure-based drug design validation. | Curated binding affinities (Ki, IC50) for model training and benchmarking. |
| GTDB & NCBI Taxonomy | Standardized microbial taxonomic classification [37] [83]. | Taxonomic dereplication & annotation. | Provides framework for clustering and interpreting genomic data. |
Table 3: Experimental Data Comparison: Performance of Representative Tools
| Tool / Method | Experimental Dataset | Key Performance Metric | Reported Outcome | Context |
|---|---|---|---|---|
| Carafe [85] | Diverse DIA proteomics datasets (global & phosphoproteome). | Peptide detection rate vs. DDA-based libraries. | Improved identification by training fragment intensity models directly on DIA data. | Spectral library generation. |
| skDER & CiDDER [37] | Enterococcus genus genomes (>1,000 genomes). | Reduction efficiency & protein space saturation. | Efficiently selected representative genomes; CiDDER achieved 95% protein cluster saturation with <20% of genomes. | Genomic dereplication. |
| Structure Landscape Analysis [87] | Integrated AFDB, ESMAtlas, and MIP databases. | Structural complementarity and functional clustering. | Databases occupy distinct but complementary regions of structure space; functions cluster in specific regions. | Database integration & analysis. |
| PredFull DNN [86] | NIST human/mouse spectral libraries with modifications. | Increase in peptide IDs after rescoring. | 8% increase in peptide IDs (21% for non-specific cleavage, 17% for phosphopeptides). | Spectral prediction for open searching. |
Experimental Protocols: From Data Generation to Validation
Protocol 1: Generating an Experiment-Specific Spectral Library with Carafe
This protocol outlines the use of Carafe, a deep learning tool that generates high-quality spectral libraries by training directly on Data-Independent Acquisition (DIA) mass spectrometry data, overcoming the mismatch with libraries generated from Data-Dependent Acquisition (DDA) data [85].
Materials:
- Software: Carafe (integrated into Skyline or standalone), DIA data analysis tool (DIA-NN or Skyline).
- Input Data: One or more DIA-MS raw files from a sample representative of the experimental system (e.g., human cell line digest). A peptide identification result file (in TSV format) from the DIA data.
Method:
- Training Data Preparation: Provide Carafe with peptide detection results (precursor m/z, charge, retention time, fragment ion intensities) from your DIA data analysis. Carafe processes this to create training data, implementing a critical step of detecting and masking "shared" fragment ion peaks arising from chimeric DIA spectra [85].
- Model Training: Fine-tune the pre-trained AlphaPeptDeep models (for fragment ion intensity and retention time prediction) using the prepared DIA training data. The training loss function excludes masked, interfered peaks to improve model accuracy [85].
- Library Generation: Use the trained model to predict fragment ion intensities and retention times for a in silico digested proteome of interest. The output is a tailored spectral library in a standard format (e.g.,
blibfor Skyline,mzSpecLib) [85]. - Validation: Apply the new library to re-analyze the DIA dataset. Compare the number of confidently identified peptides and proteins to the results obtained using a standard DDA-based or pre-trained model library.
Diagram: Carafe Spectral Library Generation Workflow
Protocol 2: Integrating Multi-Source Protein Structure Databases for Functional Analysis
This protocol describes how to create a unified structural landscape from multiple protein structure databases to explore functional complementarity, as demonstrated in recent research [87].
Materials:
- Software: Foldseek (for structural clustering), Geometricus (for structure embedding), PaCMAP (for dimensionality reduction), deepFRI (for functional annotation).
- Input Data: Representative sets from major structure databases: AlphaFold Protein Structure Database (AFDB), a high-quality subset of ESMAtlas, and the Microbiome Immunity Project (MIP) database [87].
Method:
- Dataset Curation & Redundancy Removal: Download representative structure sets from AFDB, ESMAtlas, and MIP. Perform intra-database structural clustering using Foldseek to remove redundancy within each source. Then, cluster all representatives together to remove inter-database redundancy, resulting in a non-redundant set of unique structural clusters [87].
- Structural Feature Embedding: Encode each protein structure in the unified set into a fixed-length feature vector using Geometricus, which captures 3D shape-based descriptors ("shape-mers") [87].
- Dimensionality Reduction & Mapping: Reduce the high-dimensional shape-mer vectors to a two-dimensional space using PaCMAP to create a visualizable "structural landscape" [87].
- Functional Annotation: Annotate all structures with Gene Ontology (GO) terms using a structure-based function prediction tool like deepFRI [87].
- Analysis: Visualize the 2D map. Color points by source database to assess complementarity. Color by functional annotation to identify regions of the structural landscape enriched for specific biological processes. Analyze whether novel structures from metagenomic (ESMAtlas) or focused (MIP) projects occupy distinct functional regions from well-studied proteomes (AFDB).
Diagram: Structure Database Integration and Analysis Workflow
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 4: Key Research Reagent Solutions for Database Curation and Analysis
| Category | Item / Resource | Function / Description | Example Source / Tool |
|---|---|---|---|
| Spectral Library Sources | Empirical Spectral Libraries | Collections of experimentally observed peptide spectra used for matching in proteomics searches. | NIST Libraries, PRIDE repository [86] [89]. |
| In silico Prediction Tools | Generate predicted spectra for peptides, enabling library creation for any sequence/modification. | Carafe [85], Prosit [86], PredFull [86]. | |
| Protein Structure Sources | Experimental Structure Repository | Archive of experimentally determined 3D structures (X-ray, Cryo-EM, NMR). | Protein Data Bank (PDB) [6] [84]. |
| Predicted Structure Databases | Vast collections of AI-predicted protein models for nearly all known sequences. | AlphaFold DB [87], ESMAtlas [87]. | |
| Bioactivity Data | Ligand-Target Activity Databases | Curated records of binding affinities and bioactivities for small molecules. | ChEMBL [6], BindingDB [6]. |
| Dereplication & Clustering | Genome Clustering Tools | Efficiently compute similarity (ANI) between thousands of microbial genomes. | skani (used in skDER) [37], FastANI. |
| Protein Sequence Clustering | Cluster amino acid sequences at high speed to define protein families. | CD-HIT [37], MMseqs2. | |
| Reference Curation | Taxonomic Framework | Standardized phylogenetic taxonomy for consistent classification. | Genome Taxonomy Database (GTDB) [37], NCBI Taxonomy. |
| Sequence Database Curation Workflow | Pipeline to build and filter custom reference sequence databases. | DB4Q2 workflow [83]. | |
| Integrated Analysis Platforms | Proteomics Data Analysis | Software for targeted/untargeted analysis of mass spectrometry data. | Skyline [85], DIA-NN [85], Spectronaut. |
| Structural Biology & Drug Design | Suites for molecular visualization, docking, and dynamics simulations. | Schrödinger Suite, PyMOL, OpenBabel. |
Integrated Workflows and Future Outlook
The frontier of biomolecular discovery lies in the strategic integration of taxonomic and structural approaches. A powerful sequential workflow begins with taxonomic dereplication to manage scale and bias: tools like skDER can reduce thousands of microbial genomes to a tractable set of representatives [37]. The protein complement of these representative genomes can then be funneled into structure-based analysis. Predicted structures from AFDB or ESMAtlas for key, uncharacterized proteins can be mapped onto the unified structural landscape to hypothesize function based on positional clustering with annotated proteins [87]. Subsequently, these structures can serve as targets for virtual screening in drug discovery campaigns [6] [88].
Conversely, structure-based findings can inform taxonomic studies. The discovery of a novel enzymatic function in a metagenomic protein via structural analysis [87] can trigger targeted searches for homologous sequences in genomic databases, using sensitive, structure-informed sequence profiles to expand the known taxonomic distribution of that function.
The critical enabler of this integration is the curation of high-quality, interoperable databases. Future progress depends on continued efforts to improve data quality (e.g., better peak masking in spectral libraries [85], quality metrics for predicted structures [84]), standardize annotations, and develop tools that seamlessly traverse the spectrum from sequence to structure to function. For researchers, the decision is not to choose one paradigm over the other, but to understand how the judicious application of both—leveraging their respective strengths—can provide a more complete and mechanistic understanding of biological systems.
Diagram: Integrating Taxonomic and Structure-Based Analysis Pathways
Head-to-Head Evaluation: Performance Metrics, Cost-Benefit, and Strategic Selection
The initial phase of drug discovery is defined by the strategic choice between two foundational paradigms: taxonomic-focused dereplication and structure-based design. This choice fundamentally shapes the objectives, workflows, and, ultimately, the metrics used to define success.
Taxonomic-focused dereplication is a knowledge-driven approach, primarily used in natural product (NP) discovery. It aims to quickly identify known compounds within a complex biological extract by leveraging prior knowledge organized along three pillars: the taxonomy (biological source) of the organism, the molecular structures of known metabolites, and associated spectroscopic data [31]. The primary goal is to avoid the redundant "rediscovery" of known compounds, thereby conserving resources for the pursuit of truly novel chemistry. Success in this paradigm is measured by the ability to efficiently navigate known chemical space and to identify outliers that represent new chemical entities (NCEs).
In contrast, structure-based design is a prediction-driven approach. It utilizes the three-dimensional structure of a biological target (e.g., a protein) to computationally screen, model, or design molecules that can interact with it [6]. This paradigm, which includes methods like molecular docking and free-energy calculations, seeks to rationally predict bioactivity. Its success is traditionally quantified by the accuracy of its predictions, often validated by the binding affinity (e.g., Kd, Ki) of synthesized compounds and the experimental hit rate—the proportion of tested compounds that show the desired activity.
The following diagram illustrates how these two distinct starting points lead to different primary success metrics.
Conceptual Workflow of Two Drug Discovery Paradigms
This article provides a comparative guide to the core success metrics emanating from these paradigms: Novel Compound Rate (NCR), Experimental Hit Rate (EHR), and Binding Affinity. We will define each metric, present comparative performance data from contemporary research, detail relevant experimental protocols, and discuss their strategic implications within a modern drug discovery thesis.
Comparative Analysis of Core Success Metrics
The value and interpretation of a success metric are intrinsically linked to the discovery phase and paradigm. The table below provides a foundational comparison.
Table 1: Definition and Strategic Context of Key Success Metrics
| Metric | Primary Paradigm | Definition & Calculation | Strategic Purpose | Typical Benchmark (Current) |
|---|---|---|---|---|
| Novel Compound Rate (NCR) | Taxonomic Dereplication | (Number of Novel Compounds Identified) / (Total Compounds Investigated) |
To maximize the discovery of new chemical entities and avoid redundant effort. | Highly variable; success is a non-rediscovery [31]. |
| Experimental Hit Rate (EHR) | Structure-Based Design / HTS | (Number of Confirmed Active Compounds) / (Total Compounds Tested) |
To assess the predictive accuracy of a model or the richness of a library for a given target. | HTS: ~2% [90]. AI/VS: 20-60% for top methods [90] [91]. |
| Binding Affinity (Kd, Ki) | Structure-Based Design | The concentration at which 50% of the target is bound (Kd) or inhibited (Ki). Measured via kinetics (k_off / k_on) or equilibrium assays. |
To quantify compound potency and drive structure-activity relationship (SAR) optimization. | Hit stage: µM to nM range. Lead stage: <100 nM. |
Novel Compound Rate (NCR) in Dereplication
In dereplication, a "hit" is a novel structure. The NCR is effective when supported by robust taxonomic and spectroscopic databases [31]. The process involves comparing analytical data (e.g., HR-MS, NMR) from a new sample against databases of known compounds, filtered by the taxonomic lineage of the source organism. A high-performing dereplication pipeline minimizes false negatives (mistaking a known compound for novel) and efficiently flags true unknowns for full structure elucidation.
Experimental Hit Rate (EHR) in AI & Virtual Screening
The EHR is the most direct measure of campaign efficiency in target-focused screening. Traditional high-throughput screening (HTS) of random compound libraries historically yields hit rates around 2% [90]. Modern computational methods, particularly AI-driven virtual screening, claim dramatically higher rates. However, reported EHRs must be scrutinized for the phase of discovery (e.g., Hit Identification vs. Hit Optimization) and the activity threshold used [90].
Table 2: Reported Experimental Hit Rates from AI-Driven Hit Identification Campaigns
| Model / Platform | Target | Compounds Tested | Hit Threshold | Experimental Hit Rate (EHR) | Reported Chemical Novelty (Avg. Tanimoto <0.5) |
|---|---|---|---|---|---|
| ChemPrint (Model Medicines) [90] | AXL | 29 | ≤20 µM | 41% (12/29) | Yes (0.40 vs. training/ChEMBL) |
| ChemPrint (Model Medicines) [90] | BRD4 | 12 | ≤20 µM | 58% (7/12) | Yes (0.30-0.31 vs. training/ChEMBL) |
| Schrödinger Modern VS Workflow [91] | Multiple | Varies by campaign | Not Specified | "Double-digit" hit rates | Implied by discovery of "diverse chemotypes" |
| LSTM RNN Model [90] | DRD2 | Not Specified | ≤20 µM | 43% | No (0.66 vs. training/ChEMBL) |
| Traditional HTS Benchmark [90] | General | Large Libraries | Varies | ~2% | Not Applicable |
A critical insight is that a high EHR does not guarantee novel chemistry. As shown in Table 2, some models with high EHRs (e.g., LSTM RNN) produce compounds with high similarity to known actives (Tanimoto >0.5), indicating "rediscovery" within the structure-based paradigm [90]. Therefore, the Novel Compound Rate for AI—measuring the fraction of hits that are chemically novel—is an essential complementary metric.
Binding Affinity as the Gold Standard of Potency
Binding affinity (Kd/Ki) is the definitive quantitative measure of a compound's interaction strength with its target. It is a critical endpoint for validating structure-based predictions. Accurate prediction of binding affinity remains a "grand challenge" in computational chemistry, as many scoring functions correlate poorly with experimental results [92]. Advanced methods like Free Energy Perturbation (FEP) calculations, as used in Schrödinger's FEP+, aim to bridge this gap by providing more rigorous physics-based estimates [91].
It is crucial to understand that affinity is a composite kinetic metric: Kd = koff / kon [92]. This means two compounds with identical Kd values can have very different binding kinetics (e.g., fast on/fast off vs. slow on/slow off), which can have significant implications for therapeutic efficacy and duration of action.
Experimental Protocols for Metric Validation
Protocol: Real-Time Kinetic Binding Assay for Affinity (Kd) and Kinetics (kon, koff)
This protocol, adapted from advanced cell-based studies, allows for the simultaneous determination of binding kinetics and affinity, moving beyond equilibrium measurements [93].
Objective: To determine the association (k_on) and dissociation (k_off) rate constants, and calculate the equilibrium dissociation constant (Kd = k_off / k_on) for a radiolabeled ligand.
Key Reagents & Equipment:
- Cells: Adherent cells (e.g., CHO-K1) stably expressing the target protein of interest.
- Ligand: Radiolabeled target agonist/antagonist (e.g., [¹²⁵I]-AB-MECA for the A3 adenosine receptor).
- Instrument: LigandTracer series (Ridgeview Instruments) or equivalent real-time cell-binding monitor.
- Media: Serum-free cell culture medium for assay.
Method:
- Cell Preparation: Seed cells on a section of a tilted cell culture dish to create a confluent monolayer. Incubate for 72 hours prior to assay.
- Baseline Measurement: Place the dish on the instrument detector. Add serum-free medium and record baseline radioactivity for 10-20 minutes.
- Association Phase: Introduce a known concentration of the radioligand into the medium. The instrument continuously rotates the dish, measuring the differential radioactivity between the cell-covered and cell-free areas, generating a real-time association binding curve.
- Dissociation Phase: After binding reaches near-equilibrium, remove the radioligand-containing medium and replace it with fresh ligand-free medium. Monitor the decrease in bound signal over time to generate the dissociation curve.
- Data Analysis: Fit the association curve to the equation
Y = Ymax * (1 - exp(-k_obs * t)), wherek_obsis the observed rate constant. For a single concentration of ligand (L),k_obs = k_on * (L) + k_off. By performing the assay with at least three different ligand concentrations, plotk_obsvs.(L). The slope of the line isk_onand the y-intercept isk_off. CalculateKd = k_off / k_on.
Relevance: This protocol provides a more physiologically relevant measure of affinity and crucial kinetic parameters often overlooked in standard equilibrium assays [93].
Protocol: Modern Virtual Screening Workflow for Hit Identification
This workflow describes an integrated computational-experimental pipeline to achieve high EHR with novel compounds [91].
Objective: To screen ultra-large chemical libraries (billions of compounds) to identify potent, novel hits for experimental validation.
Key Software & Resources:
- Library: Enamine REAL or similar ultra-large enumerable library.
- Docking: Glide docking software with Active Learning (AL-Glide) for machine-learning-accelerated screening.
- Rescoring: Glide WS for water-based scoring, followed by Absolute Binding Free Energy Perturbation+ (ABFEP+) for rigorous affinity prediction.
- Compound Source: Commercially available compounds for purchase or synthesis.
Method:
- Ultra-Large Library Screening:
- Pre-filter a multi-billion compound library based on physicochemical properties.
- Use AL-Glide to iteratively dock a subset of compounds, training a machine learning model to predict docking scores for the entire library, dramatically reducing computational cost.
- Perform full-precision Glide docking on the top several million compounds ranked by the ML model.
- Hierarchical Rescoring:
- Rescore the top docking hits (tens of thousands) using Glide WS, which explicitly models water molecules for improved pose prediction and enrichment.
- Select the most promising diverse chemotypes (thousands) for rigorous ABFEP+ calculations. This physics-based method provides accurate absolute binding free energy estimates.
- Experimental Triaging & Testing:
- Prioritize compounds based on predicted affinity (from ABFEP+), synthetic accessibility, and drug-like properties.
- Procure or synthesize a focused set (tens to hundreds) of top-ranked compounds.
- Test compounds in vitro using a relevant bioassay (e.g., enzymatic inhibition, cell-based activity) to determine experimental EHR and measure binding affinity (Kd/IC₅₀).
Relevance: This protocol demonstrates how integrating machine learning for scalability with physics-based methods for accuracy can transform virtual screening from a low-yield tool into a primary engine for high-EHR, high-quality hit discovery [91].
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 3: Key Reagents, Databases, and Tools for Success Metric Analysis
| Item Name | Type | Primary Function in Metric Context | Key Provider / Source |
|---|---|---|---|
| LigandTracer | Instrumentation | Enables real-time, kinetic cell-based binding assays to measure k_on, k_off, and Kd [93]. |
Ridgeview Instruments AB |
| ChEMBL Database | Bioinformatics Database | Public repository of bioactive molecules with curated binding affinities (Ki, Kd, IC₅₀). Used for model training, benchmarking, and novelty assessment [90] [6]. | EMBL-EBI |
| Glide & FEP+ | Software Suite | Industry-standard molecular docking (Glide) and high-accuracy binding free energy calculation (FEP+) platform for structure-based design [91]. | Schrödinger |
| Enamine REAL Library | Chemical Library | Ultra-large, virtually enumerated library of synthetically accessible compounds (>20B molecules) for expansive virtual screening [91]. | Enamine |
| Global Natural Products Social Molecular Networking (GNPS) | Bioinformatics Platform | Web-based mass spectrometry ecosystem for dereplication of natural products via spectral matching and molecular networking [30]. | UC San Diego |
| Tanimoto Similarity (ECFP4) | Computational Metric | Calculates molecular fingerprint similarity (0-1). Used to quantify chemical novelty of hits against training sets or known actives (novelty threshold typically <0.5) [90]. | Open-source (RDKit) |
| PDBbind Database | Bioinformatics Database | Curated database linking Protein Data Bank (PDB) structures with experimental binding affinity data. Essential for benchmarking scoring functions [6]. | PDBbind Team |
| KNApSAcK/UNPD/Coconut | NP Databases | Comprehensive databases linking natural products, their taxonomic sources, and spectra. Foundational for taxonomic dereplication [31]. | Various Academic Consortia |
Synthesis and Strategic Implications
The choice between dereplication and structure-based approaches is not merely technical but strategic, influencing resource allocation and the very definition of project success.
Integrating Metrics for a Holistic View: The most progressive discovery campaigns now seek to optimize multiple metrics simultaneously. For example, an ideal AI-driven Hit Identification campaign would score highly on three axes: a high Experimental Hit Rate (EHR), a high Novel Compound Rate (NCR) among those hits (Tanimoto <0.5 against known actives), and the subsequent confirmation of strong Binding Affinity (sub-µM) for the novel hits [90]. This moves beyond simply finding "a hit" to finding "a novel, potent hit."
Strategic Recommendations:
- For Novelty-Priority Projects (e.g., natural product discovery, pioneering new target chemotypes): Adopt a dereplication-first mindset. Use taxonomic and spectral databases aggressively to filter out known compounds. Success is measured by a high NCR and the structural novelty of the final isolates [31].
- For Potency-Priority Projects (e.g., lead optimization for a validated target): Employ structure-based design with high-accuracy affinity prediction (e.g., FEP). The primary metrics are improved Binding Affinity and maintaining/improving drug-like properties. EHR is relevant in early stages to validate the screening model.
- For Integrated AI-Driven Discovery: Demand transparent reporting of both EHR and NCR. A platform claiming a 50% EHR is only transformative if a significant subset of those hits are chemically novel and possess good affinity. Prospective validation studies should report all three metrics [90] [6].
In conclusion, "success" in modern drug discovery is multi-dimensional. Novel Compound Rate (NCR) guards against intellectual redundancy and expands chemical space. Experimental Hit Rate (EHR) measures the predictive efficiency of a chosen strategy. Binding Affinity validates the functional potency of the output. A sophisticated research thesis will not rely on a single metric but will strategically select and integrate these measures to guide the journey from hypothesis to novel therapeutic candidate.
The relentless pursuit of novel therapeutic agents demands continuous innovation in the processes that underpin drug discovery. Within this domain, natural product (NP) research remains a cornerstone for identifying unique chemotypes with potent biological activities. However, this field faces significant bottlenecks, primarily in dereplication—the early identification of known compounds to avoid redundant research—and in the subsequent resource-intensive steps of isolation and characterization [30]. These challenges frame a critical thesis in modern pharmacognosy: the strategic choice between taxonomy-focused dereplication and structure-based approaches has profound implications for both lead discovery efficiency and the optimal allocation of finite research resources.
Historically, NP discovery has been an expensive and time-consuming endeavor, with major hurdles in dereplication and structure elucidation [30]. The contemporary revival of NP studies is fueled by their value as renewable sources of medicinal compounds but is tempered by the need for greater efficiency [11]. This analysis objectively compares two principal methodological paradigms—taxonomic prioritization versus structural analysis—evaluating their performance in accelerating lead discovery while prudently managing computational, analytical, and experimental assets. The integration of artificial intelligence (AI) and high-throughput workflows further transforms this landscape, offering new pathways to reconcile depth of analysis with speed and cost-effectiveness [94] [95].
Methodological Paradigms: A Comparative Foundation
The dereplication process serves as the critical gatekeeper in NP discovery. The choice of strategy directly influences downstream resource expenditure.
Taxonomy-Focused Dereplication: This approach leverages the biological and evolutionary context of the source material. It is predicated on the understanding that taxonomically related organisms often produce structurally similar secondary metabolites. The process begins with precise taxonomic identification of the source organism. Subsequent analysis, typically via techniques like Liquid Chromatography-Mass Spectrometry (LC-MS) or Nuclear Magnetic Resonance (NMR), is guided by targeted databases containing known compounds from that specific taxon or related groups [11]. This method significantly narrows the search space, focusing resources on the most probable leads.
Structure-Based Dereplication: This paradigm prioritizes the chemical data itself, independent of biological origin. It involves the comprehensive analysis of spectroscopic and spectrometric data (e.g., MS/MS, 1D/2D NMR) from a complex mixture or purified compound. This data is then compared against vast, generic structural databases. Advances in computer-assisted structure elucidation (CASE) and tools like the Global Natural Products Social Molecular Networking (GNPS) platform exemplify this approach, enabling the identification of novel scaffolds and known compounds based purely on spectral patterns and molecular networking [30].
The decision flow for selecting the appropriate dereplication strategy is illustrated below.
Performance and Efficiency Metrics
The efficacy of dereplication strategies can be quantified through metrics related to speed, computational burden, and success rate in lead identification. The following table summarizes a comparative analysis based on current methodologies and reported data.
Table 1: Performance Comparison of Dereplication Strategies
| Metric | Taxonomy-Focused Approach | Structure-Based Approach | Key Supporting Evidence & Context |
|---|---|---|---|
| Primary Speed Advantage | Rapid preliminary filtering and annotation. | Direct, definitive structural identification when matches exist. | CNMR_Predict workflow creates searchable taxon-specific DBs for quick matching [11]. |
| Computational Resource Intensity | Lower (post-DB creation). Targeted searches require less processing. | Very High. Requires processing complex spectral data against massive DBs; AI/ML modeling adds to load [94]. | MetaflowX benchmark shows 14x faster, 38% less disk use vs. other pipelines [96]. AI models require significant compute [97]. |
| Success Rate in Novel Lead ID | Lower for novel scaffolds within a well-studied taxon. Higher for identifying known bioactive compounds. | Higher potential to identify novel structural classes, especially with MS/MS molecular networking. | GNPS enables discovery of novel analogs via spectral networking [30]. |
| Key Resource Bottleneck | Creation and curation of high-quality, taxon-specific databases with spectroscopic data. | Access to high-field NMR, high-res MS, and substantial computational power for data analysis. | CASE and quantum NMR calculations are resource-intensive [30]. |
| Integration with AI/ML | Used for predicting biogenetic pathways and compound occurrence within taxa. | Core application: predicting activity, toxicity, and de novo structure generation from spectral data [94]. | AI predicts anticancer, anti-inflammatory actions of NPs; models require large, curated datasets [94]. |
The integration of AI is reshaping both paradigms but is particularly transformative for structure-based methods. AI and machine learning models are now applied to predict biological activities, infer mechanisms of action, and prioritize candidates from vast digital libraries, moving ranked candidates into experimental validation pipelines [94]. The industry trend in 2025 shows a deeper convergence of AI with biotech operations, extending from discovery into development and manufacturing optimization [97]. For instance, AI-powered trial simulations using digital twins are beginning to reduce the need for large placebo groups, thereby conserving one of the most expensive resources in drug development: clinical trial capacity [95].
Experimental Protocols and Workflow Integration
To understand the practical implementation and resource demands, it is essential to examine representative experimental protocols for each approach.
Protocol for Taxonomy-Focused Dereplication (CNMR_Predict Workflow)
This protocol, designed for carbon-13 NMR-based dereplication, exemplifies a resource-efficient taxonomic strategy [11].
- Taxon Definition & Data Retrieval: Define the biological taxon of interest (e.g., Brassica rapa). Query the LOTUS database (or similar NP DBs like COCONUT) using the taxon as a search key to retrieve all associated chemical structures.
- Structure Curation: Process the downloaded structure files (e.g., SDF format) using cheminformatics tools (e.g., RDKit). Remove duplicates, correct tautomeric forms, and standardize valence representations to ensure compatibility with prediction software.
- Spectral Prediction: Import the curated structure list into specialized spectroscopic prediction software (e.g., ACD/Labs CNMR Predictor). Execute batch prediction of 13C NMR chemical shifts.
- Database Creation: Merge the predicted spectral data with the structural and taxonomic information to create a searchable, taxon-specific database.
- Experimental Dereplication: Analyze the crude or fractionated NP extract via 13C NMR (or LC-13C NMR). Query the experimental spectrum against the custom taxon-specific database to identify known compounds rapidly.
Protocol for Structure-Based Dereplication (GNPS/Molecular Networking)
This protocol leverages untargeted mass spectrometry and public data sharing for broad structural insight [30].
- LC-MS/MS Data Acquisition: Subject the NP extract to high-resolution liquid chromatography coupled with tandem mass spectrometry (LC-HRMS/MS). The method should fragment precursor ions to generate comprehensive MS/MS spectral data.
- Data Preprocessing: Convert raw spectral data to an open format (e.g., .mzML). Use tools like MZmine or MS-DIAL for peak picking, deisotoping, and alignment.
- Molecular Networking: Upload the processed data to the GNPS platform. Create a molecular network where MS/MS spectra are clustered based on spectral similarity (cosine score). Nodes represent ions, and edges connect spectra with high similarity, often indicating shared structural motifs.
- Database Query & Annotation: The network is automatically queried against reference spectral libraries within GNPS. Annotations are propagated within clusters, allowing for the putative identification of both known and novel analogs.
- Target Isolation: Based on network topology—such as unique clusters (potential novelty) or clusters linked to bioactive nodes—specific ions are targeted for subsequent fractionation and isolation for full structural elucidation.
The modern discovery pipeline is increasingly a hybrid, integrating multiple data streams. The following diagram synthesizes how AI-enhanced taxonomic and structural data converge to prioritize leads and guide resource allocation in a contemporary workflow.
The Scientist's Toolkit: Essential Research Reagent Solutions
The execution of these protocols relies on a suite of specialized tools and databases. The following table details key resources that constitute the modern NP researcher's toolkit.
Table 2: Key Research Reagent Solutions for NP Dereplication & Discovery
| Item Name | Type | Primary Function in Research | Relevance to Efficiency & Resource Allocation |
|---|---|---|---|
| LOTUS Database | Database | Provides curated links between NP structures and their taxonomic origins. | Enables rapid construction of taxon-specific DBs, drastically reducing manual curation time [11]. |
| GNPS Platform | Cloud Software Platform | Facilitates community-wide sharing of MS/MS spectra and automated molecular networking. | Eliminates need for in-house library generation for MS; allows annotation via crowd-sourced data, saving years of work [30]. |
| ACD/Labs CNMR Predictor | Commercial Software | Predicts 13C NMR chemical shifts for organic structures. | Replaces need for experimental reference spectra for every known compound, saving analytical time and reference materials [11]. |
| MetaflowX | Computational Workflow | Integrates reference-based and reference-free metagenomic analysis. | Benchmark shows 14x speedup and 38% less disk usage, optimizing compute resource allocation [96]. |
| CASE Software | Software Suite | Computer-Assisted Structure Elucidation uses NMR data to propose plausible structures. | Reduces time and expert labor required for the complex puzzle-solving of novel structure elucidation [30]. |
| AI/ML Models (e.g., for QSAR) | Algorithmic Tool | Predicts quantitative structure-activity relationships and biological targets. | Prioritizes the most promising leads for costly in vitro/in vivo testing, funneling resources to high-probability candidates [94] [95]. |
Synthesis and Strategic Recommendations
The comparative analysis reveals that the choice between taxonomic and structural dereplication is not binary but strategic, dependent on project goals and resource constraints.
For Resource-Constrained or Targeted Discovery: The taxonomy-focused approach offers superior efficiency. When investigating a well-defined biological source with rich prior knowledge, this method allows for the rapid elimination of known compounds with minimal analytical and computational overhead. The initial investment in building a tailored database pays dividends in streamlined workflows. This approach optimally allocates resources by focusing expensive isolation and characterization efforts only on truly novel or high-priority targets within a known chemical space.
For Novelty-Driven or Untargeted Discovery: The structure-based approach, supercharged by AI and molecular networking, is indispensable. When exploring uncharted taxonomic territory or seeking entirely novel scaffolds, this paradigm casts the widest net. While computationally intensive, platforms like GNPS democratize access to powerful comparative analytics. The resource allocation here shifts from manual curation to computational power and advanced analytical instrumentation (high-res MS, high-field NMR). The return on investment is the higher potential for groundbreaking discoveries.
The prevailing trend is toward hybridization and AI integration. The most efficient future pipelines will likely start with a taxonomic filter to quickly remove common knowns, followed by a deep structural analysis of the remaining "unknowns" using AI-powered tools to predict activity and novelty. As seen in 2025 trends, AI's role is expanding from pure discovery to optimizing entire development pipelines, including clinical trial design and manufacturing, representing the ultimate strategic allocation of digital resources to conserve physical and financial assets [97] [95]. Consequently, the most effective resource allocation strategy invests in the computational infrastructure and data science expertise required to harness these converging methodologies, ensuring that every experimental dollar is guided by the maximum possible informational insight.
The discovery of novel bioactive compounds from natural sources is a cornerstone of drug development but presents a fundamental strategic dilemma. Researchers must balance the pursuit of novel chemical scaffolds against the need to understand precise target engagement and mechanism of action. This tension frames a broader methodological thesis in natural product research: taxonomy-focused dereplication versus structure-based approaches [30] [22]. Dereplication prioritizes the rapid identification and elimination of known compounds within complex extracts to spotlight novel chemical entities [30]. In contrast, structure-based approaches focus on elucidating the three-dimensional configuration of a molecule to predict and validate its interaction with a biological target, which is critical for understanding efficacy and toxicity [30].
This guide objectively compares these two paradigms, providing experimental data and protocols to inform strategic decisions in early-stage drug discovery. The optimal path is not a binary choice but a question of priority and integration, guided by project goals, resource availability, and the nature of the biological target.
Comparison of Strategic Approaches
The following table outlines the core objectives, typical workflows, and key outputs of the two primary strategies.
Table 1: Strategic Comparison of Dereplication-First vs. Structure-First Approaches
| Aspect | Dereplication-First Strategy (Novelty-Driven) | Structure-First Strategy (Target/Mechanism-Driven) |
|---|---|---|
| Primary Goal | Maximize the discovery rate of novel chemical scaffolds from natural source libraries [30] [98]. | Understand and optimize the binding affinity, specificity, and mechanism of action of a lead compound [30]. |
| Core Methodology | High-throughput analytical profiling (LC-MS/MS, Molecular Networking) combined with database searches to filter out known compounds [22] [34]. | Determination of absolute stereochemistry, computational docking, and biophysical assays (SPR, ITC, X-ray crystallography) to study target-ligand interactions [30]. |
| Ideal Application Context | Unexplored or biodiverse taxonomic sources (e.g., marine microbes, extremophiles); projects aimed at expanding chemical diversity libraries [30] [98]. | Projects with a well-defined, druggable biological target; lead optimization phases; repurposing known scaffolds for new targets [30]. |
| Key Bottlenecks | Quality and comprehensiveness of spectral and structural databases; ionization bias in MS; "silent" biosynthetic gene clusters not expressed under lab conditions [22] [98]. | Difficulty in determining absolute configuration of complex metabolites; high protein/compound requirements for structural biology methods; limited predictability of in vivo activity [30]. |
| Major Output | A prioritized list of extract fractions or pure compounds with a high probability of containing novel chemical entities [34]. | A high-resolution 3D model of the ligand-target complex, informing structure-activity relationships (SAR) and rational design [30]. |
Quantitative Performance Metrics
The choice between strategies is further clarified by their historical and operational performance metrics.
Table 2: Quantitative Performance and Yield Metrics
| Metric | Dereplication-First Approach | Structure-First/Target-Based Approach | Notes & Data Source |
|---|---|---|---|
| Throughput (samples/week) | High (100-1000+) [98]. Automated LC-MS/MS with molecular networking can process hundreds of extracts. | Low to Medium (1-10). Structure elucidation, especially absolute configuration determination, is rate-limiting [30]. | Throughput is a key differentiator in the early discovery phase. |
| Material Requirement | Low (ng-µg). Sufficient for LC-MS and MS/MS profiling [22]. | High (mg). Typically required for NMR-based structure elucidation and crystallography [22]. | Micro-cryoprobe NMR and MS reduce but do not eliminate this gap [22]. |
| Novel Scaffold Hit Rate | Higher (in novel taxa). Focused on filtering out knowns, directly increasing the odds of novelty. Reported success in expressing "silent" gene clusters [98]. | Variable/Lower. May rediscover known scaffolds that are novel binders for a specific target. | Hit rate is highly dependent on the pre-screening biological model and source diversity. |
| Success Rate to Pre-clinical Candidate | Lower. Novel scaffold does not guarantee drug-like properties or tolerable toxicity. | Higher (for validated targets). Understanding target engagement de-risks downstream optimization. | Analysis of drug discovery pipelines shows target-based strategies have a higher clinical transition rate [14]. |
| Database Dependency | Critical. Relies on extensive, high-quality MS/MS and NMR spectral libraries (e.g., GNPS) [34]. | Moderate. Relies on protein data bank (PDB) and chemical structure databases. | Gaps in dereplication databases are a major source of re-discovery [22] [34]. |
Decision Framework and Integrated Workflow
The decision to prioritize dereplication or structure-based analysis is not mutually exclusive. The following diagram conceptualizes the dynamic interplay between these strategies within a modern integrated drug discovery pipeline.
Strategic Decision Points:
- Prioritize Dereplication When: The project is in the exploratory phase with access to unique or underexplored taxonomic sources (e.g., marine or extremophile microorganisms) [98]. The goal is to build a library of novel chemotypes for broad phenotypic screening or to address a high risk of rediscovery. This is particularly powerful when guided by genome mining that predicts silent biosynthetic gene clusters (BGCs) [98].
- Prioritize Structure-Based Analysis When: The project is target-driven, with a clear, validated protein target implicated in a disease. The goal is to find a potent and selective binder, which may involve repurposing a known scaffold. This approach is critical for solving mechanism of action and enabling rational medicinal chemistry optimization [30].
- Integrate Both Approaches: The most robust strategy involves iteration. Novel scaffolds from dereplication must progress to target engagement studies. Conversely, known compounds identified in target-based screens should be dereplicated immediately to avoid wasted effort. Molecular networking is a key integrative tool, visually organizing related compounds and annotating structures directly from complex mixtures [34].
Experimental Protocols
Protocol 1: High-Throughput Dereplication via LC-MS/MS and Molecular Networking
This protocol is designed for the rapid prioritization of extracts containing novel natural products [22] [34].
Sample Preparation:
- Prepare microbial or plant extracts in a suitable solvent (e.g., methanol, ethyl acetate). For microbial strains, consider using HiTES (High-Throughput Elicitor Screening) to activate silent BGCs under hundreds of culture conditions prior to extraction [98].
- Use a 96-well or 384-well plate format. Include blank solvent controls and, if available, internal standards.
LC-MS/MS Analysis:
- Instrument: Employ a UHPLC system coupled to a high-resolution tandem mass spectrometer (e.g., Q-TOF, Orbitrap).
- Chromatography: Use a reversed-phase C18 column with a standard water-acetonitrile gradient. Keep run times short (5-15 minutes) for high throughput.
- Mass Spectrometry: Operate in data-dependent acquisition (DDA) mode. Acquire full-scan MS1 spectra (e.g., m/z 100-1500) at high resolution (>30,000). Select the top N most intense ions from each scan for fragmentation (MS2) using collision-induced dissociation (CID).
Data Processing and Molecular Networking (via GNPS):
- Convert raw data to open formats (.mzML, .mzXML) using tools like MSConvert.
- Upload data to the Global Natural Products Social Molecular Networking (GNPS) platform [34].
- Create a Feature-Based Molecular Network (FBMN). This workflow uses software like MZmine3 to first detect chromatographic features (aligning m/z, retention time, and intensity) before networking, improving accuracy [34].
- Set parameters: minimum cosine score for spectral similarity (e.g., 0.7), minimum matched fragment ions (e.g., 6). The network will cluster molecules with similar MS2 spectra, implying structural relatedness.
- Perform dereplication by searching MS2 spectra against public (e.g., GNPS libraries) and in-house spectral libraries. Compounds with high-score matches are considered "known."
Prioritization:
- Prioritize for further isolation: Singleton nodes (not connected to any known compound cluster) or clusters containing no annotated nodes.
- Also prioritize clusters where known compounds are connected to many unannotated "daughter" nodes, suggesting novel derivatives of a known scaffold [34].
Protocol 2: Determining Absolute Configuration for Target Engagement Studies
This protocol outlines steps to unambiguously determine the 3D structure of a novel active compound, a prerequisite for meaningful docking studies and SAR [30].
Purification and Preliminary Data:
- Isolve the active compound to high purity (>95%) using preparative HPLC.
- Acquire standard 1D and 2D NMR data (¹H, ¹³C, COSY, HSQC, HMBC) to establish the planar structure.
Computational Chemistry Predictions:
- Generate all possible stereoisomers of the planar structure.
- For each isomer, perform conformational searches and geometry optimization using quantum chemical methods (e.g., Density Functional Theory - DFT).
- Calculate the theoretical NMR parameters (chemical shifts, coupling constants) and optical rotation (OR) or electronic circular dichroism (ECD) spectra for the lowest-energy conformers of each isomer.
- Tools: Use tools like Gaussian or ORCA for calculations, and DP4 probability analysis to statistically compare calculated vs. experimental NMR shifts [30].
Experimental Stereochemical Analysis:
- Experimental Chiroptical Data: Acquire experimental OR and ECD spectra of the purified compound.
- Advanced NMR: Acquire NOESY or ROESY spectra to obtain through-space proton-proton correlations, which provide relative configuration data.
- Chemical Derivatization (Mosher's Method): For compounds with a single stereogenic center bearing a hydroxyl or amine group, synthesize (R)- and (S)-Mosher ester derivatives. The Δδ (δS – δR) values of nearby protons in the ¹H NMR spectra can determine absolute configuration.
Structure-Target Docking:
- Prepare the 3D structure of the ligand with the determined absolute configuration.
- Obtain the 3D structure of the target protein (from PDB or homology modeling).
- Perform molecular docking simulations (e.g., using AutoDock Vina, Glide) to predict binding poses, affinity, and key interactions (hydrogen bonds, hydrophobic contacts).
- Validation: If resources allow, validate the docking pose by solving the co-crystal structure of the ligand bound to the target protein via X-ray crystallography.
Research Reagent Solutions Toolkit
Table 3: Essential Research Tools and Reagents
| Category | Item/Technique | Primary Function in Discovery Pipeline | Key Consideration |
|---|---|---|---|
| Dereplication & Analytics | High-Resolution LC-MS/MS (Q-TOF, Orbitrap) | Provides accurate mass (for formula prediction) and MS/MS spectra (for structural similarity networking) of compounds in complex mixtures [22] [34]. | High mass accuracy (<5 ppm) and fast scanning speeds are critical for high-throughput profiling. |
| Global Natural Products Social Molecular Networking (GNPS) | An online platform for processing MS/MS data to create visual molecular networks, enabling dereplication and novelty detection [34]. | Open-access and community-driven; requires data in specific open formats (.mzML). | |
| Feature-Based Molecular Networking (FBMN) | An advanced GNPS workflow that incorporates chromatographic peak alignment, improving network reliability for complex samples [34]. | Requires additional upstream processing with tools like MZmine3 or OpenMS. | |
| Structure Elucidation | NMR Spectroscopy (500 MHz+) | Determines covalent connectivity (planar structure) and, via NOESY/ROESY, relative configuration of pure compounds [30] [22]. | Milligram quantities needed; micro-cryoprobes enhance sensitivity. The major bottleneck in structure elucidation. |
| Computer-Assisted Structure Elucidation (CASE) Software | Uses algorithms to generate all possible structures consistent with experimental NMR and MS data, ranking them by probability [30]. | Reduces time for solving complex planar structures but still requires expert interpretation. | |
| Quantum Chemistry Software (e.g., Gaussian) | Calculates theoretical NMR, OR, and ECD spectra for candidate stereoisomers to compare with experimental data and assign absolute configuration [30]. | Computationally intensive; requires expertise in computational chemistry. | |
| Target Engagement | Surface Plasmon Resonance (SPR) | A label-free technique to measure real-time binding kinetics (kon, koff) and affinity (KD) between a compound and an immobilized protein target. | Provides direct evidence of binding; requires purified, functional protein. |
| Differential Scanning Fluorimetry (Thermal Shift Assay) | Measures protein thermal stabilization upon ligand binding, indicating direct target engagement in a low-cost, medium-throughput format. | Excellent for initial screening of fragment or compound libraries against a purified target. | |
| Molecular Docking Software (e.g., AutoDock, Glide) | Predicts the preferred orientation (pose) and binding affinity of a small molecule within a protein's active site. | A computational tool for hypothesis generation; poses must be validated experimentally. | |
| Enabling Technologies | Genome Mining Tools (e.g., antiSMASH) | Identifies biosynthetic gene clusters (BGCs) in microbial genomes, predicting chemical potential and guiding strain prioritization [98]. | Essential for a taxonomy-focused, genome-driven discovery strategy. |
| Heterologous Expression Systems | Expresses silent BGCs from unculturable or pathogenic microbes in tractable model hosts (e.g., Aspergillus nidulans, S. albus) for compound production [98]. | Solves supply and regulatory issues for compounds from difficult sources. |
The process of early drug discovery is fundamentally an exercise in intelligent prioritization. With ultra-large chemical libraries now containing billions of purchasable and make-on-demand compounds, the central challenge has shifted from mere access to chemical space to the efficient navigation of it [99] [100]. Virtual screening (VS) serves as a primary compass, yet it generates an overwhelming number of putative hits, many of which are false positives, known compounds, or synthetically intractable. This is where the synergistic integration of dereplication—the fast identification of known compounds—into the VS pipeline becomes a critical strategic advantage [11].
This integration must be understood within the context of a broader methodological thesis: the comparison between taxonomy-focused and structure-based dereplication paradigms. Taxonomy-focused dereplication, grounded in natural product research, leverages biological context (e.g., the species, genus, or family of a source organism) to constrain the identification process. It operates on the principle of taxonomic relatedness, where organisms produce structurally related secondary metabolites [11]. In contrast, structure-based dereplication, more common in synthetic library screening, relies purely on physico-chemical data—such as molecular fingerprints, spectral matches (MS, NMR), or predicted properties—to filter out rediscovered chemotypes [11].
The synergy proposed here uses dereplication not as a post-screening checkpoint, but as an integrated filter that prunes virtual hits before they enter the synthesis queue. This guides medicinal chemists toward novel, tractable, and potent leads, directly addressing the critical "Make" bottleneck in the Design-Make-Test-Analyse (DMTA) cycle [100]. By framing this workflow within the aforementioned thesis, we can objectively compare how the biological logic of taxonomy and the physical logic of computational structure prediction can be combined to create a more robust and efficient discovery engine.
Methodology for Comparative Performance Analysis
Experimental Protocols for Benchmarking Virtual Screening Tools
The performance of structure-based virtual screening (SBVS) tools, a key component of the pipeline, was evaluated using established benchmarking protocols. A standardized method involves using the DEKOIS 2.0 benchmark sets, which provide known active molecules and carefully generated decoy molecules for specific protein targets [101]. The performance is assessed by a docking tool's ability to rank active molecules above decoys.
Typical Protocol [101]:
- Protein Preparation: Crystal structures (e.g., from PDB) are prepared by removing water molecules and co-crystallized ligands, adding hydrogen atoms, and optimizing side-chain conformations using tools like UCSF Chimera or the OpenEye toolkit.
- Ligand/Decoy Preparation: Active ligands and decoys from the DEKOIS set are prepared: generating tautomers, protonation states at physiological pH, and multiple conformations using software like Omega2 or RDKit.
- Docking Execution: Prepared ligands are docked into the defined binding site of the prepared protein using various docking software (e.g., AutoDock Vina, PLANTS, FRED). A standardized grid box ensures consistent search space.
- Performance Metrics: The primary metric is Enrichment Factor at 1% (EF1%), measuring how many actives are found in the top 1% of the ranked list compared to a random distribution. Other metrics include the area under the Receiver Operating Characteristic curve (ROC-AUC) and pROC-Chemotype plots to assess chemotype diversity among top-ranked hits.
- Re-scoring with Machine Learning (ML): Docking poses are re-evaluated using pretrained ML scoring functions (e.g., CNN-Score, RF-Score-VS v2) to test if ML can improve the prioritization of true actives [101].
Experimental Protocol for Taxonomy-Focused Dereplication
For taxonomy-focused dereplication, especially relevant in natural product-based screening, a key protocol involves creating and using taxon-specific databases [11].
Typical Protocol [11]:
- Taxon Definition & Database Query: The biological source of the extract or the target taxonomic group is defined (e.g., Brassica rapa). A comprehensive natural product database like LOTUS (which links structures to taxonomic data) is queried using the taxon name.
- Database Curation & Enhancement: The resulting list of candidate structures is downloaded and curated (removing duplicates, standardizing tautomeric forms). To enable spectroscopic dereplication, predicted 13C NMR chemical shifts are computationally added for all candidates using specialized prediction software (e.g., ACD/Labs CNMR Predictor).
- Dereplication Matching: Experimental data (e.g., HR-MS for molecular formula, 13C NMR shifts from a purified fraction) are compared against the curated, taxon-focused database. A match within a defined tolerance filter (e.g., ± 0.5 ppm for NMR) provides a high-confidence identification, flagging the compound as "known" for that taxon.
Integrated Workflow Protocol
A synergistic workflow integrates the above components sequentially, as demonstrated in a study identifying PARP-1 inhibitors [102].
- Ultra-Library Pre-filtering: An ultra-large library (e.g., ~13 million molecules) is first filtered for drug-likeness (e.g., Lipinski's Rule of Five) and undesirable substructures.
- AI-Powered Ligand-Based Screening: A deep learning model (e.g., TransFoxMol, a graph neural network combined with a Transformer) is used to score and rank the filtered library based on predicted activity against the target [102].
- Structure-Based Docking & Consensus Ranking: The top-ranked molecules from the AI model (e.g., top 100,000) are subjected to docking with one or more docking tools (e.g., KarmaDock, AutoDock Vina). Poses and scores are generated, and a consensus ranking is created.
- Integrated Dereplication Filter: The top virtual hits (e.g., top 1,000) are subjected to dereplication. This involves:
- Structural Dereplication: Checking against major databases of known bioactive compounds (e.g., ChEMBL, PubChem) via molecular fingerprint similarity (Tanimoto coefficient).
- Taxonomic Dereplication (if applicable): If the screen is based on a natural product-inspired library, checks against taxon-specific databases are performed.
- Synthetic Tractability Filter: Proposed hits are analyzed for synthetic accessibility using AI retrosynthesis tools (e.g., AIZynthFinder) or rule-based scoring (SA Score) [100].
- Final Prioritization & Synthesis Guidance: The compounds surviving the dereplication and tractability filters are clustered by scaffold. Representatives from promising clusters are selected for synthesis, guided by predicted routes from Computer-Assisted Synthesis Planning (CASP) tools [100].
Results & Discussion: Comparative Performance of Integrated Strategies
Performance of Standalone vs. Integrated Dereplication Strategies
The table below summarizes the core characteristics and performance implications of taxonomy-focused and structure-based dereplication when used in isolation versus in an integrated model.
Table 1: Comparison of Dereplication Strategies and Their Role in an Integrated Workflow
| Strategy | Core Principle | Primary Data Input | Key Advantage | Major Limitation | Role in Integrated VS Pipeline |
|---|---|---|---|---|---|
| Taxonomy-Focused Dereplication [11] | Biological relatedness predicts chemical similarity. | Taxon of biological source; Experimental/Predicted NMR or MS spectra. | Drastically reduces candidate space; High confidence in identification within a taxon. | Limited to natural products; Requires well-defined taxonomy; Database coverage gaps. | Early filter for natural product libraries; Prevents re-isolation of known metabolites from related species. |
| Structure-Based Dereplication | Physicochemical similarity indicates identity. | 2D/3D molecular structure; Spectroscopic fingerprints. | Broadly applicable to any compound; Amenable to high-throughput computational screening. | Can miss known compounds with different representations; Prone to false negatives with novel scaffolds. | Post-docking filter to remove known bioactives and frequent hitters from synthetic libraries. |
| Synergistic Integration (This Work) | Serial application of biological and physicochemical logic. | Combined taxonomic, structural, and synthetic data. | Maximizes novelty and tractability of hits; Bridges biological context with computational prediction. | Increased workflow complexity; Requires multi-disciplinary data integration. | Central workflow controller: Guides synthesis by filtering VS hits through successive lenses of novelty (dereplication) and feasibility (synthesis planning). |
Benchmarking Virtual Screening Tools for the Structure-Based Component
The accuracy of the SBVS component is critical. A comprehensive 2025 benchmark evaluated multiple docking methods across key dimensions, revealing a clear performance hierarchy [103].
Table 2: Performance Benchmark of Docking Methods in Pose Prediction and Physical Validity (Summarized from [103])
| Method Category | Example Tools | Pose Accuracy (RMSD ≤ 2Å) | Physical Validity (PB-Valid Rate) | Combined Success Rate | Key Finding |
|---|---|---|---|---|---|
| Traditional Physics-Based | Glide SP, AutoDock Vina | Moderate to High | Very High (≥94%) | High | Excel in producing physically plausible poses; robust across diverse targets. |
| Generative Diffusion Models | SurfDock, DiffBindFR | Very High (≥70%) | Moderate to Low | Moderate | Superior pose accuracy but often generate physically implausible interactions (clashes, bad angles). |
| Regression-Based Models | KarmaDock, GAABind | Low | Very Low | Low | Often fail to produce valid molecular geometries despite predicting affinity. |
| Hybrid Methods | Interformer | High | High | Highest | Best balance, combining AI scoring with traditional conformational search for reliable, valid poses. |
Furthermore, re-scoring docking outputs with Machine Learning Scoring Functions (ML-SFs) significantly enriches hit rates. A benchmark on antimalarial target PfDHFR showed that re-scoring with CNN-Score boosted early enrichment (EF1%) from worse-than-random to as high as 31 for a drug-resistant variant [101]. This demonstrates that an integrated VS workflow using a traditional or hybrid docking tool followed by ML re-scoring is a high-performance strategy for the structure-based component.
Quantitative Impact of Dereplication on Synthesis Guidance
The ultimate goal of integration is to funnel resources toward synthesizing the most promising novel hits. A quantitative model of SBVS performance underscores this, showing that hit rates plateau and can even drop at the very top of a ranked list due to scoring artifacts [104]. This model emphasizes that physically testing compounds across a range of ranks is essential to find the true peak hit-rate. Dereplication directly addresses this by removing such artifacts (often known promiscuous binders) and known compounds, effectively "cleaning" the top of the list and increasing the probability that synthesized compounds will be novel and genuine hits.
The synthesis planning ("Make") step is the primary bottleneck [100]. By applying dereplication and synthetic accessibility scoring before synthesis, the integrated workflow ensures that medicinal chemistry efforts are focused. AI-powered synthesis planning tools can then generate routes for the final, vetted list of novel virtual hits, dramatically accelerating the cycle. For instance, platforms like Exscientia report AI-driven design cycles that are ~70% faster and require 10x fewer synthesized compounds than industry norms [26].
Visualizing the Integrated Workflow and Strategic Thesis
The following diagrams, created using Graphviz DOT language, illustrate the integrated workflow and the conceptual thesis framing the research.
Synergistic VS Workflow with Integrated Dereplication Filter
Thesis Framework: Integrating Dereplication Paradigms
Table 3: Key Reagent Solutions for Implementing the Integrated Workflow
| Tool/Resource Category | Specific Examples | Primary Function in the Workflow | Key Reference / Source |
|---|---|---|---|
| Natural Product / Taxonomy Databases | LOTUS, COCONUT, KNApSAcK | Provides the structural and taxonomic data essential for taxonomy-focused dereplication; links compounds to biological sources. | [11] |
| Spectroscopic Prediction Software | ACD/Labs CNMR Predictor, NMRShiftDB | Predicts NMR (or MS) spectra for database compounds, enabling spectral matching for dereplication without isolated standards. | [11] |
| Ultra-Large Chemical Libraries | Enamine REAL, Topscience Database, WuXi MADE | Provides the source chemical space (billions of compounds) for virtual screening. "Make-on-Demand" libraries vastly expand accessible novelty. | [102] [100] |
| AI Ligand-Based VS Models | TransFoxMol, Graph Neural Network (GNN) models | Performs initial, rapid scoring of ultra-large libraries based on learned structure-activity relationships, enabling a tractable pre-filter for docking. | [102] |
| Molecular Docking Software | AutoDock Vina, PLANTS, FRED, KarmaDock | Performs structure-based virtual screening by predicting binding poses and generating initial affinity scores for protein-ligand complexes. | [103] [102] [101] |
| Machine Learning Scoring Functions | CNN-Score, RF-Score-VS v2 | Re-scores and re-ranks docking outputs to significantly improve the enrichment of true active compounds over decoys. | [101] |
| Cheminformatics Toolkits | RDKit, Open Babel | Provides fundamental capabilities for molecule manipulation, descriptor calculation, fingerprint generation, and file format conversion throughout the pipeline. | [11] [102] |
| Synthesis Planning & Tractability | AIZynthFinder, CASP tools, SA Score | Evaluates the synthetic accessibility of virtual hits and proposes potential retrosynthetic routes, guiding the final "Make" decision. | [100] |
| Building Block Sourcing Platforms | Enamine, eMolecules, MolPort | Provides access to physical and virtual building blocks for the synthesis of prioritized hits, integrated via inventory management systems. | [100] |
The field of natural product (NP) discovery and drug development is undergoing a transformative shift, driven by the convergence of two powerful paradigms: AI-powered predictive modeling and integrated hybrid discovery platforms. This evolution is fundamentally reshaping the long-standing methodological debate between taxonomy-focused dereplication and structure-based approaches [11] [22]. Taxonomy-focused methods prioritize the biological origin of compounds, leveraging phylogenetic relationships to narrow chemical search spaces [11]. In contrast, structure-based approaches use spectroscopic data, such as mass spectrometry (MS) or nuclear magnetic resonance (NMR), to identify compounds directly from complex mixtures without primary reliance on biological activity or source [22]. The integration of AI models capable of predicting molecular properties, binding affinities, and even de novo designs with automated, data-connected laboratory platforms is creating a new, synergistic workflow. This convergence promises to overcome the individual limitations of each classical approach, accelerating the path from biological material to validated lead compounds [105] [106] [107].
Comparative Analysis of Dereplication Approaches and AI Integration
The choice between taxonomic and structure-based dereplication involves trade-offs between specificity, sensitivity, and throughput. The integration of AI and automation is enhancing the capabilities of both.
Table 1: Core Comparison of Dereplication Methodologies
| Feature | Taxonomy-Focused Dereplication | Structure-Based Dereplication | Role of AI/Hybrid Convergence |
|---|---|---|---|
| Primary Driver | Biological origin & phylogenetic relationship [11]. | Spectroscopic/spectrometric data of the compound [22]. | Unification: AI models integrate taxonomic priors with structural data for higher-confidence annotation. |
| Typical Data | Taxonomic databases (e.g., LOTUS), 13C NMR predicted shifts [11]. | MS/MS fragmentation patterns, 1H/13C NMR spectra [22]. | Multi-modal Learning: AI (e.g., foundation models) fuses MS, NMR, and genomic data for holistic identification [105] [106]. |
| Key Advantage | Reduces candidate pool using evolutionary constraints; high relevance for known taxa [11]. | Can discover novel scaffolds unrelated to known bioactivity; high-throughput via LC-MS [22]. | Predictive Power: AI predicts NMR/ MS spectra and bioactivity from structure, bridging identification and function [107]. |
| Main Limitation | Misses compounds from horizontal gene transfer or new taxa; depends on database completeness [11]. | Can generate "unknowable" compounds with no known activity; requires pure compounds for NMR [22]. | Automated Workflows: Hybrid platforms automate from sample prep to data analysis, feeding AI with clean, structured data [105]. |
| Automation & AI Readiness | Medium. Requires curated taxonomic-structure databases. AI can predict taxon-specific chemical space [11]. | High. MS data is inherently digital and high-throughput. Ideal for AI-powered spectral matching and de novo interpretation [106] [22]. | Platform Integration: Solutions like Cenevo and Sonrai Analytics connect data, instruments, and AI to close the loop from experiment to insight [105]. |
Table 2: Performance Metrics of AI-Powered Predictive Models in Drug Discovery (2025 Analysis)
| AI Model / Platform | Primary Function | Reported Performance / Impact | Experimental Validation |
|---|---|---|---|
| Boltz-2 (MIT/Recursion) | Predict protein-ligand binding affinity [107]. | Top predictor at CASP16; calculates affinity 1,000x faster than physics-based FEP simulations [107]. | Validated on curated datasets from ChEMBL and BindingDB; powers the SAIR repository of 5.2 million computed structures [107]. |
| Hermes (Leash Bio) | Predict small molecule-protein binding likelihood [107]. | 200-500x faster than Boltz-2 with improved accuracy on proprietary benchmarks; simple architecture (sequence & SMILES input) [107]. | Trained on large, high-quality proprietary dataset to minimize "batch effect" noise; enables hit expansion via Artemis tool [107]. |
| Latent-X (Latent Labs) | De novo design of therapeutic proteins (mini-binders, macrocycles) [107]. | Achieves picomolar binding affinity testing only 30-100 candidates per target (vs. millions in HTS) [107]. | Head-to-head experimental comparisons show competitive binding vs. state-of-the-art (RFdiffusion, AlphaProteo) [107]. |
| AI in Systematic Review (2025) | Various applications across drug development [106]. | 40.9% of studies used ML; 39.3% of AI applications were in preclinical stage; 72.8% focused on oncology [106]. | Analysis of 173 studies shows AI enhances drug efficacy and trial outcomes; 97% of studies reported industry partnerships [106]. |
| Foundation Models (e.g., Sonrai Analytics) | Multi-modal data integration (imaging, omics, clinical) [105]. | Extracts features from histopathology slides to identify novel biomarkers and link to clinical outcomes [105]. | Applied in trusted research environments with transparent workflows to build regulatory and partner trust [105]. |
Detailed Experimental Protocols
The convergence of AI and hybrid platforms is operationalized through novel, streamlined experimental protocols.
Protocol 1: Creating a Taxonomy-Focused 13C NMR Database with AI-Enhanced Prediction This protocol, adapted from contemporary methods, details the creation of a targeted dereplication database [11].
- Objective: To build a taxon-specific database of natural products with predicted 13C NMR shifts for rapid dereplication.
- Materials:
- Method:
- Compound Retrieval: Query the LOTUS database using the taxonomic name of the organism under study (e.g., Brassica rapa). Export the resulting list of associated chemical structures in SDF format [11].
- Data Curation: Process the SDF file using Python scripts to remove duplicates (via InChI keys), correct tautomeric forms (e.g., convert iminols to amides), and standardize valence representations for compatibility with prediction software [11].
- AI-Enhanced Prediction: Import the curated SDF file into the prediction software. Execute batch 13C NMR chemical shift prediction. Modern AI-based predictors significantly accelerate this previously bottlenecked step and can offer higher accuracy for complex or novel scaffolds [11] [107].
- Database Deployment: Export the final dataset, which now contains structures, taxonomic information, and predicted spectra, into a searchable format (e.g., a local SQL database or integration into a platform like GNPS). This database can now be used to query experimental 13C NMR data from purified compounds or even partially resolved mixtures [11].
Protocol 2: Validating AI-Predicted Binding Affinity with Hybrid Screening Platforms This protocol validates computational hits from models like Boltz-2 or Hermes in a biologically relevant context [105] [107].
- Objective: To experimentally test the binding affinity and functional activity of small molecules selected by an AI prediction model.
- Materials:
- AI Predictions: List of candidate molecules and their predicted binding scores/affinities from a model (e.g., Boltz-2 output) [107].
- Hybrid Discovery Platform: An integrated system like Nuclera's eProtein Discovery System for rapid protein expression or mo:re's MO:BOT for automated 3D cell culture [105].
- Assay Reagents: Purified target protein (from Protocol 2.1 or commercial source) or human-relevant cell lines (from Protocol 2.2).
- Method:
- Target Production (if needed): Use an automated protein expression and purification system (e.g., Nuclera's platform) to produce the soluble, active target protein for binding assays. This step can be completed in under 48 hours, overcoming a traditional bottleneck [105].
- Biological Model Preparation: Seed and maintain human-relevant cell models (e.g., organoids in a 96-well format) using an automated cell culture platform (e.g., MO:BOT). This ensures standardized, reproducible biological material for phenotypic screening [105].
- Experimental Screening:
- For biochemical assays: Perform a binding assay (e.g., fluorescence polarization, surface plasmon resonance) using the purified protein and the AI-selected compounds.
- For phenotypic assays: Treat the standardized organoids with the compounds and use automated, high-content imaging to read out phenotypic changes.
- Data Integration & Model Feedback: Feed the experimental binding or activity data back into the AI model. This closed-loop process, managed by data integration platforms like Cenevo or Labguru, retrains and improves the predictive algorithm for subsequent cycles [105].
Visualizing the Converged Workflow
The following diagram illustrates the synergistic workflow created by the convergence of AI models and hybrid platforms, bridging taxonomy and structure-based approaches.
AI-Hybrid Platform Convergence Workflow
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 3: Key Research Reagents & Platforms for Convergent Discovery
| Item / Solution | Function in Workflow | Relevance to Thesis Context |
|---|---|---|
| LOTUS Database [11] | Provides the critical link between chemical structures and the taxonomy of the organisms that produce them. | Core to taxonomy-focused dereplication. Enables creation of taxon-specific search databases. |
| ACD/Labs CNMR Predictor or AI-based Alternatives [11] | Predicts 13C NMR chemical shifts for organic structures, populating databases when experimental data is missing. | Accelerates the bottleneck in building taxonomy-focused NMR databases; modern AI versions increase accuracy. |
| Global Natural Products Social (GNPS) Molecular Networking [22] | A crowdsourced platform for MS/MS spectral data sharing and dereplication via molecular networking. | Core to structure-based approach. Allows unknown MS/MS spectra to be compared against a vast community database. |
| SAIR (Structurally-Augmented IC50 Repository) [107] | An open-access repository of computationally folded protein-ligand structures with experimental affinity data. | Trains & validates AI binding prediction models like Boltz-2, bridging structural prediction and experimental activity. |
| RDKit Cheminformatics Library [11] | An open-source toolkit for cheminformatics used to manipulate chemical structures, handle SDF files, and calculate descriptors. | Essential for curating and standardizing chemical structure data from various sources before AI analysis. |
| Automated 3D Cell Culture Platform (e.g., MO:BOT) [105] | Standardizes the production of human-relevant tissue models (organoids) for phenotypic screening. | Provides biologically relevant validation for candidates from either approach, enhancing translation potential. |
| Integrated Data Platform (e.g., Labguru, Cenevo) [105] | Connects instruments, manages experiments, and structures metadata to create AI-ready datasets. | The "central nervous system" of the hybrid platform, ensuring traceability and feeding clean data to AI models. |
| ChEMBL / BindingDB [107] | Public databases containing bioactive molecules with drug-like properties, and binding affinities. | Primary sources of experimental data for training and benchmarking AI models for binding and activity prediction. |
Future Directions and Strategic Recommendations
The trajectory points towards deeper integration and more autonomous discovery cycles. Key future directions include:
- The Rise of Self-Driving Laboratories: The full integration of AI decision-making with robotic liquid handlers, automated analyzers, and seamless data platforms will enable closed-loop "design-make-test-analyze" cycles with minimal human intervention. Platforms like Tecan's Veya and SPT Labtech's firefly+ exemplify the move toward such integrated, flexible automation [105].
- Physics-Informed AI Models: Combining the generalizability of deep learning with the rigorous constraints of physical laws (as seen in SandboxAQ's LQMs) will yield models that are both accurate for prediction and trustworthy in interpretation, especially for critical parameters like binding affinity [107].
- Explainable AI (XAI) for Regulatory Science: As AI-designed candidates move toward the clinic, transparency in decision-making is paramount. Platforms like Sonrai Analytics emphasize completely open workflows to build trust with regulators and partners [105].
- Democratization Through Open Tools: The release of powerful models like Boltz-2 under permissive licenses and the growth of open repositories like SAIR lower barriers to entry, allowing academia and smaller biotechs to leverage state-of-the-art predictive tools [107].
Strategic Recommendation: Research teams should adopt a hybridized strategy. Begin with a taxonomy-focused screen to leverage evolutionary wisdom and quickly isolate known bioactive compounds. Subsequently, apply structure-based AI models to the remaining "unknown" fractions to discover novel scaffolds. This sequential approach, powered by an integrated data and automation platform, maximizes the efficiency and success rate of natural product discovery and drug development programs.
Conclusion
Taxonomic-focused dereplication and structure-based approaches are not mutually exclusive but represent complementary axes in the modern drug discovery landscape. Dereplication excels as a high-throughput filter to navigate chemical space and safeguard against rediscovery, proving indispensable in natural product research and microbiome analysis[citation:2][citation:3][citation:9]. Structure-based methods provide a mechanistic, target-driven framework for rational design and activity prediction, increasingly augmented by AI and sophisticated simulations[citation:1][citation:6]. The optimal strategy is context-dependent, dictated by project goals, available data, and resource constraints. The future lies in integrated platforms that seamlessly combine the prioritization power of dereplication with the predictive, mechanism-based insights of structural modeling. This synergy, fueled by advances in cheminformatics, machine learning, and multi-omics, will accelerate the discovery of novel, efficacious therapeutics for complex diseases[citation:4][citation:7][citation:10].
References
- 1. Rapid Dereplication of Bioactive Compounds in Plant and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12380011/]
- 2. An effective dereplication strategy for analyzing a ... [https://www.sciencedirect.com/science/article/abs/pii/S0026265X25018867]
- 3. Comparing alignment and de-novo approaches for gut ... [https://www.nature.com/articles/s41598-025-26617-6]
- 4. An integrated multi-omic approach for discovery and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12560952/]
- 5. A globally integrated structure of taxonomy to support ... [https://www.sciencedirect.com/science/article/pii/S016953472300215X]
- 6. Navigating structure-based drug discovery with emerging ... [https://www.nature.com/articles/s44386-025-00031-4]
- 7. Structure-Based Drug Design Summit: Home [https://structure-based-drug-design-summit.com/]
- 8. Dereplication of microbial metabolites through database ... [https://www.nature.com/articles/s41467-018-06082-8]
- 9. Structure-Based Drug Design Conference 2025 - Schrödinger [https://www.schrodinger.com/life-science/resources/event/structure-based-drug-design-conference-2025/]
- 10. Modern Approaches in the Discovery and Development of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8779633/]
- 11. Taxonomy-Focused Natural Product Databases for Carbon ... [https://www.mdpi.com/2673-4532/2/3/6]
- 12. A Structure-Based Drug Discovery Paradigm - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC6601033/]
- 13. Structure and dynamics in drug discovery [https://www.nature.com/articles/s44386-024-00001-2]
- 14. Natural products in drug discovery: advances and ... [https://www.nature.com/articles/s41573-020-00114-z]
- 15. Fragment- and structure-based drug discovery for developing ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8666131/]
- 16. Dereplication Based Strategy for Rapid Identification and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8600527/]
- 17. Development of a High-Resolution Tandem Mass Spectral ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12537953/]
- 18. Metagenomic Analysis Using Phylogenetic Placement—A ... [https://www.frontiersin.org/journals/bioinformatics/articles/10.3389/fbinf.2022.871393/full]
- 19. MaLiAmPi enables generalizable and taxonomy ... [https://www.sciencedirect.com/science/article/pii/S2667237523003107]
- 20. Comparing alignment and de-novo approaches for gut ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12660995/]
- 21. Integrating taxonomic signals from MAGs and contigs ... [https://www.nature.com/articles/s41467-024-47155-1]
- 22. Modern mass spectrometry for synthetic biology and structure ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4981503/]
- 23. Enhancing drug discovery through structure-based design ... [https://www.frontiersin.org/journals/drug-discovery/articles/10.3389/fddsv.2025.1632015/full]
- 24. Difference between Molecular Docking and Molecular ... [https://www.iaanalysis.com/molecular-dynamics-simulation-docking-difference.html]
- 25. Dynamic Docking: A Paradigm Shift in Computational Drug ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6150405/]
- 26. Leading AI-Driven Drug Discovery Platforms [https://www.sciencedirect.com/science/article/abs/pii/S0031699725075118]
- 27. DynamicBind: predicting ligand-specific protein ... [https://www.nature.com/articles/s41467-024-45461-2]
- 28. NMR-driven structure-based drug discovery by unveiling ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12126535/]
- 29. Structure based screening and molecular docking with ... [https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0307615]
- 30. Advanced Methods for Natural Products Discovery: Bioactivity ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10222211/]
- 31. The Three Pillars of Natural Product Dereplication. Alkaloids ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7865595/]
- 32. Integrated genome mining and molecular networking ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12366066/]
- 33. Molecular Networking in Cosmetic Analysis: A Review of Non ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12526295/]
- 34. Molecular networking: An efficient tool for discovering and ... [https://www.sciencedirect.com/science/article/abs/pii/S0731708525000822]
- 35. Non-targeted metabolomics-based molecular networking ... [https://link.springer.com/article/10.1007/s11306-024-02210-2]
- 36. Automated genome mining predicts structural diversity and ... [https://elifesciences.org/reviewed-preprints/109154]
- 37. skDER & CiDDER: two scalable approaches for microbial ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10690176/]
- 38. GNPS - Analyze, Connect, and Network with your Mass ... [https://gnps.ucsd.edu/]
- 39. Homology modeling, molecular docking and MD ... [https://pubmed.ncbi.nlm.nih.gov/40192075/]
- 40. "Molecular Dynamics Simulations and Reverse Docking ... [https://www.sciencedirect.com/science/article/pii/S1872204025001252]
- 41. Benchmarking the Structure-Based Virtual Screening ... [https://pubmed.ncbi.nlm.nih.gov/40837276/]
- 42. A comparative study of computational modeling ... [https://www.nature.com/articles/s41598-025-16866-w]
- 43. State of the Art of Protein Structure Prediction as of 2025 ... [https://medium.com/advances-in-biological-science/state-of-the-art-of-protein-structure-prediction-as-of-2025-as-evaluated-by-casp16-0c423636bc97]
- 44. DeepFold-PLM: accelerating protein structure prediction ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12598648/]
- 45. Comparison of software for molecular mechanics modeling [https://en.wikipedia.org/wiki/Comparison_of_software_for_molecular_mechanics_modeling]
- 46. Best CPU, GPU, RAM for Molecular Dynamics in 2024 ... [https://bizon-tech.com/blog/best-cpu-gpu-ram-for-molecular-dynamics?srsltid=AfmBOoow4keIkUA4i8sd-QQy6jeD8lAbsWmtuUkGqr2UUAYT_C_Za2rR]
- 47. A Comparative Analysis of Cheminformatics Platforms [https://intuitionlabs.ai/articles/cheminformatics-platforms-comparison]
- 48. Top 9 Applications of Cheminformatics in the Biopharma ... [https://neovarsity.org/blogs/cheminformatics-applications-biopharma-2025]
- 49. Scalability of mass spectrometry-based metabolomics for ... [https://pubs.rsc.org/en/content/articlehtml/2025/np/d5np00034c]
- 50. AI-Integrated QSAR Modeling for Enhanced Drug Discovery [https://pmc.ncbi.nlm.nih.gov/articles/PMC12525248/]
- 51. Cheminformatics Software [https://www.simulations-plus.com/software/cheminformatics-software/]
- 52. Top Drug Discovery Software Solutions to Watch in 2025 [https://www.deepmirror.ai/post/top-drug-discovery-software-solutions-to-watch-in-2025]
- 53. Federated Learning for ADMET Prediction [https://www.apheris.com/resources/blog/federated-learning-for-admet-prediction-expanding-model-applicability]
- 54. Benchmarking ML in ADMET predictions: the practical impact ... [https://jcheminf.biomedcentral.com/articles/10.1186/s13321-025-01041-0]
- 55. A new genomic blueprint of the human gut microbiota [https://www.nature.com/articles/s41586-019-0965-1]
- 56. Genome-resolved metagenomics: a game changer for ... [https://www.nature.com/articles/s12276-024-01262-7]
- 57. Discovery of Reactive Microbiota-Derived Metabolites that ... [https://www.sciencedirect.com/science/article/pii/S0092867416317378]
- 58. Comparing genomes recovered from time-series ... - Microbiome [https://microbiomejournal.biomedcentral.com/articles/10.1186/s40168-023-01557-3]
- 59. MAGdb: a comprehensive high quality MAGs repository for ... [https://genomebiology.biomedcentral.com/articles/10.1186/s13059-025-03711-6]
- 60. Computational protein-ligand docking and virtual drug ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4868550/]
- 61. Identification of potential natural BCL-2 inhibitors for leukemia ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12570718/]
- 62. Identification of natural inhibitors against CDK6 target ... [https://www.sciencedirect.com/science/article/abs/pii/S1476927125001744]
- 63. Discovery of a novel potent tubulin inhibitor through virtual ... [https://www.nature.com/articles/s41420-025-02679-3]
- 64. Applications of the Pharmacophore Concept in Natural ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7685156/]
- 65. A pharmacophore-guided deep learning approach for ... [https://www.nature.com/articles/s41467-023-41454-9]
- 66. Generation of appropriate protein structures for virtual ... [https://www.sciencedirect.com/science/article/pii/S2950363925000286]
- 67. Interformer: an interaction-aware model for protein-ligand ... [https://www.nature.com/articles/s41467-024-54440-6]
- 68. Metagenomic insights into the taxonomy, function, and ... [https://microbiomejournal.biomedcentral.com/articles/10.1186/s40168-021-01031-y]
- 69. Insights into secondary metabolism from a global analysis ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4123684/]
- 70. Metabolomics and Genomics in Natural Products Research [https://pmc.ncbi.nlm.nih.gov/articles/PMC8691422/]
- 71. Comparative mapping of sequence-based and structure ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC1087832/]
- 72. Docking strategies for predicting protein-ligand interactions ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11583305/]
- 73. Structure prediction of protein-ligand complexes from ... [https://www.nature.com/articles/s41467-024-48837-6]
- 74. Data as a Product for Structure-Based Drug Design (SBDD) [https://www.desertsci.com/2025/04/17/data-as-a-product-for-structure-based-drug-design-sbdd/]
- 75. FBMN Workflow - GNPS Documentation - GitHub Pages [https://ccms-ucsd.github.io/GNPSDocumentation/featurebasedmolecularnetworking/]
- 76. Ensemble docking for intrinsically disordered proteins - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC11785235/]
- 77. An artificial intelligence accelerated virtual screening ... [https://www.nature.com/articles/s41467-024-52061-7]
- 78. Harnessing free energy calculations for kinome-wide ... [https://www.nature.com/articles/s41467-025-62722-w]
- 79. Pose ensemble graph neural networks to improve docking ... [https://pubs.rsc.org/en/content/articlehtml/2025/sc/d4sc07875f]
- 80. Enhancing multifunctional drug screening via artificial ... [https://pubs.rsc.org/en/content/articlehtml/2025/dd/d5dd00082c]
- 81. Quality over quantity: how to get the best results when ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12146287/]
- 82. AI meets physics in computational structure-based drug ... [https://www.nature.com/articles/s44386-025-00019-0]
- 83. A detailed workflow to develop QIIME2-formatted reference databases for taxonomic analysis of DNA metabarcoding data | BMC Genomic Data | Full Text [https://bmcgenomdata.biomedcentral.com/articles/10.1186/s12863-022-01067-5]
- 84. current challenges in structure-based drug design [https://www.sciencedirect.com/science/article/abs/pii/S1359644624002319]
- 85. Carafe enables high quality in silico spectral library ... [https://www.nature.com/articles/s41467-025-64928-4]
- 86. exploring more of the proteome using mass spectral libraries ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10542943/]
- 87. Large protein databases reveal structural complementarity ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12378989/]
- 88. Leveraging ligand-based and structure-based approaches ... [https://www.iptonline.com/articles/leveraging-ligand-based-and-structure-based-approaches-for-advan]
- 89. Top 6 Proteomics Databases and How to Access Them [https://bigomics.ch/blog/guide-to-top-proteomics-databases-and-how-to-access-them/]
- 90. AI Hit Rates and Novelty [https://modelmedicines.com/newsroom/ai-hit-rate]
- 91. Dramatically improving hit rates with a modern virtual ... [https://www.schrodinger.com/life-science/learn/white-papers/dramatically-improving-hit-rates-modern-virtual-screening-workflow/]
- 92. Binding Affinity Determination in Drug Design - PubMed Central [https://pmc.ncbi.nlm.nih.gov/articles/PMC11240957/]
- 93. New approaches for the reliable in vitro assessment of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5340791/]
- 94. Artificial intelligence for natural product drug discovery and ... [https://www.sciencedirect.com/science/article/pii/S2666521225001206]
- 95. The latest trends in drug discovery [https://www.cas.org/resources/cas-insights/2025-drug-discovery-trends]
- 96. MetaflowX: a scalable and resource-efficient workflow for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12489473/]
- 97. Five Trends That Got Everyone Talking in Pharma and ... [https://www.biospace.com/drug-development/five-trends-that-got-everyone-talking-in-pharma-and-biotech-this-year]
- 98. High-Throughput Mining of Novel Compounds from Known ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11243205/]
- 99. Combined usage of ligand- and structure-based virtual ... [https://www.sciencedirect.com/science/article/abs/pii/S0223523424010444]
- 100. Accelerating compound synthesis in drug discovery [https://pmc.ncbi.nlm.nih.gov/articles/PMC12560346/]
- 101. Benchmarking the Structure-Based Virtual Screening ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12363558/]
- 102. Development of an Integrated Computational Pipeline for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12368897/]
- 103. Decoding the limits of deep learning in molecular docking for ... [https://pubs.rsc.org/en/content/articlehtml/2025/sc/d5sc05395a]
- 104. A quantitative model of structure-based virtual screening ... [https://chemrxiv.org/engage/chemrxiv/article-details/69037e08113cc7cfff050553]
- 105. Inside ELRIG's Drug Discovery 2025 [https://www.drugtargetreview.com/article/190237/inside-elrigs-drug-discovery-2025/]
- 106. From Lab to Clinic: How Artificial Intelligence (AI) Is ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12298131/]
- 107. AI Models Enhance the Drug Discovery Portfolio [https://www.genengnews.com/topics/artificial-intelligence/ai-models-enhance-the-drug-discovery-portfolio/]